 Open Access
Open Access
ARTICLE
Exploring the utility of a NGS multigene panel to predict BCG response in patients with non-muscle invasive bladder cancer
1 Department of Molecular Medicine, Sapienza University of Rome, Rome, 00161, Italy
2 Department of Maternal-Child and Urological Sciences, Sapienza University of Rome, Rome, 00161, Italy
3 Department of Mechanical and Aerospace Engineering, Sapienza University of Rome, Rome, 00161, Italy
4 Department of Radiological, Oncological and Pathological Sciences, Sapienza University of Rome, Rome, 00161, Italy
5 Department of Urology, Stanford University School of Medicine, Stanford, CA 94305, USA
6 University Center of Excellence in Urology, Wrocław Medical University, Wrocław, 50556, Poland
7 Istituto Pasteur-Fondazione Cenci Bolognetti, Rome, 00161, Italy
* Corresponding Author: CHIARA NICOLAZZO. Email:
# These authors contributed equally to this work
Oncology Research 2025, 33(3), 723-731. https://doi.org/10.32604/or.2024.056282
Received 18 July 2024; Accepted 12 October 2024; Issue published 28 February 2025
Abstract
Objectives: Intravesical Bacillus Calmette-Guérin (BCG) therapy is a gold standard for patients with high-risk non-muscle invasive bladder cancer (NMIBC). Although a long-lasting therapeutic response is observed in most patients, BCG failure occurs in 30%–50% of patients and a progression to muscle-invasive disease is found in 10%–15%. Therefore, predicting high-risk patients who might not benefit from BCG treatment is critical. The purpose of this study was to identify, whether the presence of specific oncogenic mutations might be indicative of BCG treatment response. Methods: Nineteen high-grade NMIBC patients who received intravesical BCG were retrospectively enrolled and divided into “responders” and “non-responders” groups. Tissue samples from transurethral resection of bladder cancer were performed before starting therapy and were examined using a multigene sequencing panel. Results: Mutations in TP53, FGFR3, PIK3CA, KRAS, CTNNB1, ALK and DDR2 genes were detected. TP53 and FGFR3 were found to be the most frequently mutated genes in our cohort (31.6% and 26.3%, respectively), followed by PIK3CA (15.8%). In the BCG-responsive patient group, 90% of samples were found to have mutated genes, with almost 50% of them showing mutations in tyrosine kinase receptors and CTNNB1 genes. On the other hand, in the BCG-unresponsive group, we found mutations in 44.4% of samples, mainly in TP53 gene. Conclusions: Our findings suggest that a Next-Generation Sequencing (NGS) multigene panel is useful in predicting BCG response in patients with NMIBC.Graphic Abstract
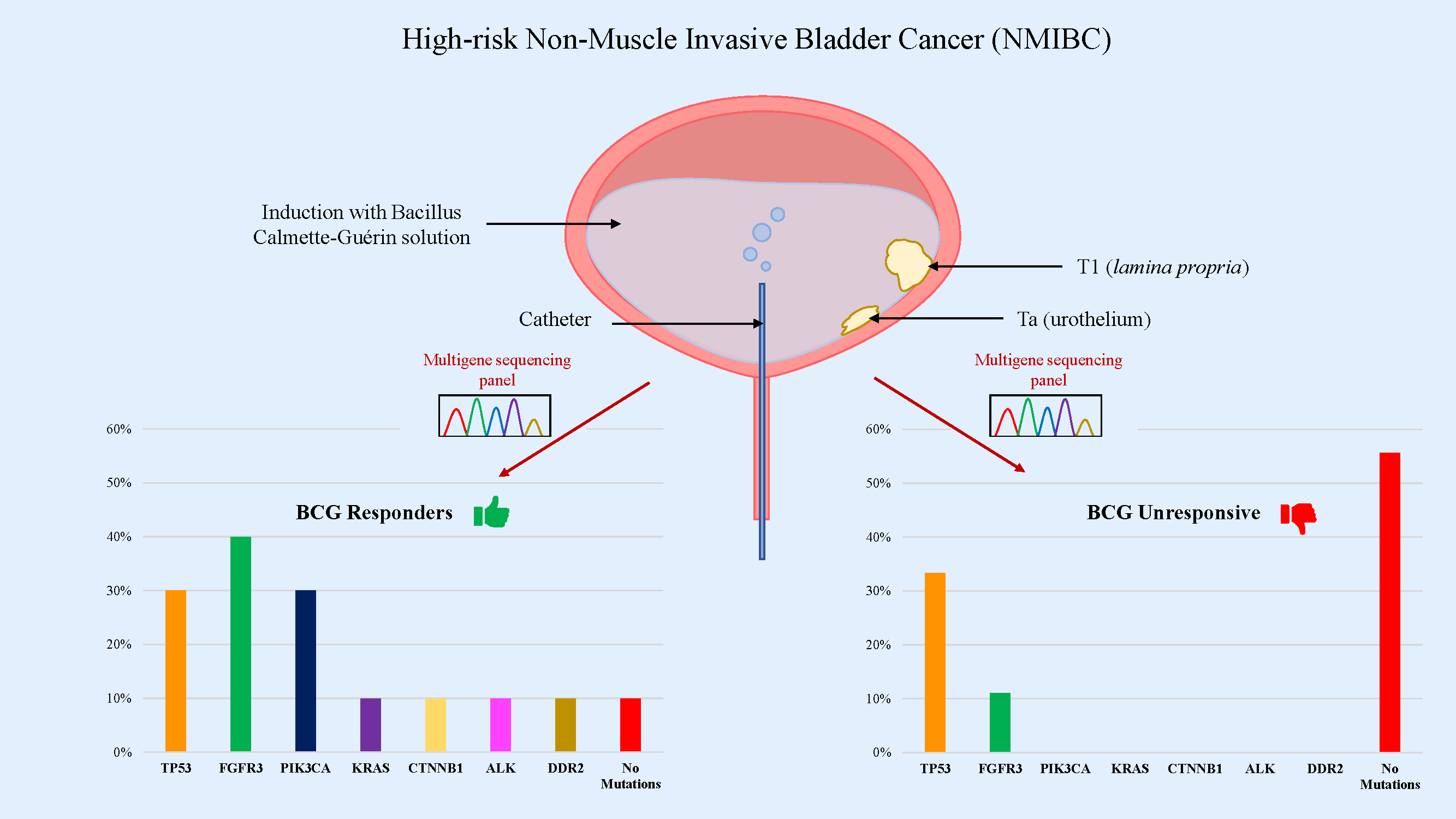
Keywords
Supplementary Material
Supplementary Material FileCite This Article
 Copyright © 2025 The Author(s). Published by Tech Science Press.
Copyright © 2025 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools