 Open Access
Open Access
ARTICLE
Leveraging Deep Learning for Precise Chronic Bronchitis Identification in X-Ray Modalities
1 School of Computing, Faculty of Technology, University of Portsmouth, Winston Churchill Ave Southsea, Portsmouth, PO1 2UP, UK
2 Portsmouth Artificial Intelligence and Data Science Center, University of Portsmouth, Winston Churchill Ave, Southsea, Portsmouth, PO1 2UP, UK
3 Department of Computer Science, College of Computer and Information Sciences, Jouf University, Sakaka, Aljouf, 72341, Saudi Arabia
4 School of Biological and Behavioural Sciences, Queen Mary University of London, London, E1 4NS, UK
5 Department of Computer Sciences, Bahria University, Lahore, Punjab, 54700, Pakistan
* Corresponding Author: Fahad Ahmad. Email:
Computers, Materials & Continua 2025, 83(1), 381-405. https://doi.org/10.32604/cmc.2025.062452
Received 18 December 2024; Accepted 10 February 2025; Issue published 26 March 2025
Abstract
Image processing plays a vital role in various fields such as autonomous systems, healthcare, and cataloging, especially when integrated with deep learning (DL). It is crucial in medical diagnostics, including the early detection of diseases like chronic obstructive pulmonary disease (COPD), which claimed 3.2 million lives in 2015. COPD, a life-threatening condition often caused by prolonged exposure to lung irritants and smoking, progresses through stages. Early diagnosis through image processing can significantly improve survival rates. COPD encompasses chronic bronchitis (CB) and emphysema; CB particularly increases in smokers and generally affects individuals between 50 and 70 years old. It damages the lungs’ air sacs, reducing oxygen transport and causing symptoms like coughing and shortness of breath. Treatments such as beta-agonists and inhaled steroids are used to manage symptoms and prolong lung function. Moreover, COVID-19 poses an additional risk to individuals with CB due to its impact on the respiratory system. The proposed system utilizes convolutional neural networks (CNN) to diagnose CB. In this system, CNN extracts essential and significant features from X-ray modalities, which are then fed into the neural network. The network undergoes training to recognize patterns and make accurate predictions based on the learned features. By leveraging DL techniques, the system aims to enhance the precision and reliability of CB detection. Our research specifically focuses on a subset of 189 lung disease images, carefully selected for model evaluation. To further refine the training process, various data augmentation and noise removal techniques are implemented. These techniques significantly enhance the quality of the training data, improving the model’s robustness and generalizability. As a result, the diagnostic accuracy has improved from 98.6% to 99.2%. This advancement not only validates the efficacy of our proposed model but also represents a significant improvement over existing literature. It highlights the potential of CNN-based approaches in transforming medical diagnostics through refined image analysis, learning capabilities, and automated feature extraction.Keywords
The human respiratory system is critically anchored by the lungs, situated just below the rib cage and above the diaphragm. These essential organs are responsible for the vital exchange of gases, absorbing oxygen into the bloodstream and expelling carbon dioxide. This exchange is integral not only for respiratory function but also for maintaining the body’s pH balance [1].
The lungs are equipped with various defence mechanisms to protect against infections. The respiratory tract is lined with cilia, hair-like projections that rhythmically transport mucus, trapping and removing dust and bacteria from inhaled air towards the pharynx. Additionally, the lung linings secrete immunoglobulins that bolster defence against respiratory pathogens [2]. Mucus produced by goblet cells contains antimicrobial agents such as defensins, antiproteases, and antioxidants, which safeguard internal organs from severe diseases [3].
Despite these defences, various factors, including genetic predispositions and environmental exposures, can precipitate lung disorders. Diseases affecting the cardiothoracic and respiratory tract regions, such as asthma, disrupt normal lung function by inflaming and narrowing airways. Lung cancer, another serious condition, manifests with symptoms like coughing up blood, chest pain, and unexplained weight loss. Often, these symptoms are nonspecific, leading to delayed detection and advanced disease stages [4].
Chronic obstructive pulmonary disease (COPD), encompassing chronic bronchitis (CB) and emphysema, is particularly debilitating. It is characterized by excessive mucus production and breathing difficulties. Lung infections, including pneumonia and bronchitis, are frequently triggered by viruses but can also be caused by bacteria and fungi. Pulmonary arterial hypertension (PAH) further complicates lung health by constricting blood vessels and increasing arterial pressure, forcing the heart to work harder and potentially leading to heart failure [5].
Environmental factors, such as allergens like pollens, mold, and air pollution, are known triggers for asthma. Smoking remains the leading risk factor for lung cancer, accounting for approximately 80% of related deaths. Early detection of these conditions is challenging as their symptoms often mimic less severe illnesses [6]. COPD, primarily induced by prolonged exposure to irritants like cigarette smoke and combustion fumes, is another major concern. Additionally, diseases like cirrhosis and autoimmune disorders can precipitate PAH [7–9].
COPD represents a significant public health challenge. Ranked as the fourth leading cause of death in the U.S., it affects approximately 11 million individuals annually. This condition combines diseases like CB, leading to persistent inflammation and deterioration of lung function. Often undiagnosed until severe symptoms emerge, COPD complicates treatment and management efforts. Common symptoms include breathlessness, recurring chest infections, and persistent wheezing. Prolonged exposure to noxious fumes, especially from smoking, increases the risk of COPD and its complications, such as respiratory infections, cardiovascular diseases, lung cancer, and mental health issues [10].
CB and emphysema, the primary forms of COPD, result from the inflammation of bronchial tubes and damage to lung air sacs, respectively. Inflamed cilia—tiny hair-like structures responsible for expelling mucus—impair mucus clearance, allowing harmful pathogens easier access to the lungs. This condition progresses as bronchial tubes swell and accumulate mucus, further disrupting the lungs’ gas exchange capabilities, thus reducing oxygen intake and carbon dioxide expulsion [11,12].
CB significantly impacts the quality of life through symptoms like excessive mucus production and persistent coughing. In the U.S., CB affects around 9 million adults, with smokers being 40% more susceptible than non-smokers. Other risk factors include prolonged exposure to air pollutants, occupational hazards, and genetic predispositions [13]. The interconnected damage caused by CB can often lead to emphysema, where alveoli walls are destroyed, creating larger air spaces and reducing oxygen transfer to the blood. This leads to oxygen deprivation in the bloodstream, particularly during physical activity.
Moreover, CB poses heightened risks during pandemics such as COVID-19. The overlapping symptoms of these conditions complicate diagnoses and exacerbate the vulnerability of already compromised lung function. Identifying the stages of CB—mild, moderate, severe, and very severe—is essential for effective management. Although not curative, treatment aims to alleviate symptoms and prevent further complications. Long-term medications, including beta-agonists, inhaled steroids, and anticholinergics, are recommended to ease symptoms and improve lung health [14].
Technological advancements in medical imaging have significantly improved the diagnosis and management of lung diseases like CB. High-resolution computed tomography (CT) scans provide detailed views of the lungs, aiding early detection and accurate assessment of disease progression. Pulmonary function tests further evaluate the impact on lung capacity and functionality [15].
In parallel, deep learning (DL) technologies have emerged as transformative tools in medical diagnostics, particularly through image analysis. DL models, especially convolutional neural networks (CNN), are increasingly used to classify and detect CB from lung images [16]. These models leverage multiple processing layers to enhance pattern recognition in complex image data, outperforming traditional diagnostic models. The integration of CNN into medical diagnostics highlights DL’s potential to differentiate between healthy and affected lung tissues, offering a critical tool in combating CB and related pulmonary conditions [17]. The seamless integration of DL into medical imaging enriches clinical assessments while ensuring a connected, data-driven approach to healthcare. By automating the detection and classification of lung conditions from images, DL facilitates quicker, more accurate diagnoses, ultimately improving patient outcomes and streamlining healthcare processes.
In response to the identified need for improved diagnostic methods for CB, our study introduces a CNN based approach that overcomes the limitations of traditional diagnostic techniques. Existing methods often lack the precision and efficiency required for early and accurate detection of CB, primarily due to inadequate feature extraction and image analysis capabilities. To address the scarcity of available datasets, our approach incorporates data augmentation to enhance the dataset size, alongside advanced pre-processing techniques for more effective feature extraction. These improvements allow CNN to achieve better diagnostic results by identifying subtle pathological features often overlooked by conventional methods.
This research initiative is cantered on utilizing DL to enhance the precision of CB identification in X-ray modalities. The study aims to develop and refine a CNN based methodology for diagnosing CB, employing augmented X-ray datasets to overcome challenges related to limited data availability. The core challenge involves leveraging data augmentation techniques to expand dataset size effectively, enabling the DL model to perform accurate and reliable CB diagnosis.
The primary aim of this study is to expand the dataset size using augmentation techniques, thereby enhancing the realism and precision of CB identification through advanced DL in X-ray modalities. We strive to improve upon existing classification systems in terms of efficacy. The objective extends beyond simply refining the analytical methods; it also aims to equip medical practitioners with a more accurate diagnostic tool. This enhancement will support better-informed decision-making in CB treatment, which is particularly useful when existing datasets are limited.
The key contribution of this study lies in its systematic approach to enlarging the dataset through various augmentation techniques and assessing the efficacy of the CNN algorithm in classifying CB. By enhancing the dataset and fine-tuning the CNN model, the study provides significant insights into the capabilities of DL technologies in medical diagnostics. This advancement not only demonstrates the potential of augmented datasets in improving diagnostic accuracy but also highlights the role of innovative DL techniques in advancing medical research and practice.
The evolving landscape of medical technology, particularly through the application of DL in image-based diagnostics, offers promising avenues for addressing the complexities of chronic lung diseases. As these technologies advance, they bring hope for more effective management and better prognostic capabilities for conditions like CB, fundamentally changing the approach to respiratory healthcare.
The rest of the paper is arranged as below. Section 2 presents the literature review, Section 3 explains the used methodology, Section 4 elaborates on the proposed approach & experimental results, Section 5 elucidates the discussion, and Section 6 concludes the paper.
This section reviews related works on DL oriented lung disease classification, emphasizing advancements in methodologies to enhance detection accuracy. Various DL techniques have been proposed to address challenges in lung disease identification and improve diagnostic outcomes.
The adaptive crisp active contour approach (ACACA) has shown effectiveness in segmenting lung structures, using co-occurrence statistic model-based features for lung disease identification. This method integrates spatial association frameworks to improve image data analysis, enabling precise differentiation between healthy and diseased lungs [18]. A contrasting study targeting the Chinese population employed machine learning (ML) algorithms, including XGBoost (Extreme Gradient Boosting), logistic regression, and K-nearest neighbours (KNN), to classify COPD, with KNN achieving the highest accuracy of 82% [19,20].
Further exploration of DL techniques led to a neural network ensemble (NNE) approach, which differentiated benign from malignant lung nodules, achieving an accuracy of 78.7%—a significant improvement over individual classifiers [21]. Similarly, the LUNA-16 challenge, initiated in 2011, leveraged raw CT scans to predict lung nodules by localizing their occurrence and assessing their likelihood within specified regions [22,23]. Building on this, a multiscale two-layer CNN applied to the Lung Image Database Consortium (LIDC) database achieved a remarkable diagnostic accuracy of 86.84% for lung cancer, highlighting the potential of hierarchical learning frameworks in medical imaging [24,25].
In cancer treatment, a study explored chemoradiotherapy combined with radiation for locally advanced non-small cell lung cancer, using neural networks to identify cancer cells [26]. Another study introduced a hybrid framework combining CNNs, deep belief networks (DBNs), and stacked denoising autoencoders (SDAEs) for cancer diagnosis, leveraging multiple image features [27,28]. Innovative research presented the first end-to-end DL-based model for CB diagnosis, utilizing CT images to identify five distinct lung tissue patterns indicative of obstructive and restrictive lung diseases, validated using two publicly available datasets [29].
An algorithm based on DL was developed to classify CB according to Fleischner Society criteria, validating its outcomes across different cohorts and correlating classifications with impaired pulmonary function tests [30]. Another study utilized X-ray imaging to develop a regression convolutional network capable of predicting CB percentages. Initial accuracy of 85.39% increased to 90.73% after model adjustments, demonstrating robust predictive capabilities for CB [31].
Additionally, a novel diagnostic system for COPD was introduced, utilizing parametric resonance map image analysis to differentiate distinct COPD types via CT scans, showcasing the utility of image-based disease characterization [32]. Another study addressed COPD classification using a multiple instance learning approach, effectively classifying CT image patches and integrating them for comprehensive diagnostic assessments, even in weakly labelled datasets [33]. A five-step methodology for diagnosing COPD through 3D CT images was proposed, segmenting lung regions, assessing parenchymal variations, and classifying disease severity through combined classification and statistical evaluation techniques [34].
Transfer learning (TL) has also been explored for chest CT image classification, demonstrating high accuracy in detecting conditions such as tuberculosis, pneumonia, and COVID-19. Using a hybrid model with angular rotations during training, this approach underscored TL’s potential for improving detection rates [35]. Diverse DL methodologies, including capsule networks, visual geometry group-oriented neural networks (VGGNNs), vanilla neural networks (VNNs), and CNNs, have been employed to address challenges in detecting lung diseases. These methods address the limitations of traditional CNNs, particularly for abnormally oriented or rotated images. For instance, a hybrid model combining CNNs, spatial transformer networks (STNs), VGGNNs, and data augmentation achieved 73% accuracy [36]. Another study used CNNs to predict lung infection severity by correlating radiological findings with ML predictions, demonstrating TL’s effectiveness in enhancing accuracy for COVID-19 severity classification [37,38].
DL techniques have consistently shown promise in detecting lung diseases from chest X-rays. Approaches such as CNNs, VGG-based networks, and hybrid models have demonstrated superior accuracy compared to traditional ML techniques [35,39]. Multi-modal strategies combining X-ray images with electronic health records have further advanced COPD detection [40]. While DL models have achieved high performance in detecting abnormalities such as lung nodules, pneumonia, and tuberculosis, challenges persist in ensuring model fairness across demographic groups and integrating these technologies into clinical practice. Despite these challenges, DL-based methods hold significant potential as screening tools for lung diseases, particularly in resource-limited settings where access to specialized diagnostic equipment is restricted [41].
Below is a comparative Table 1 summarizing the different studies and their key findings:
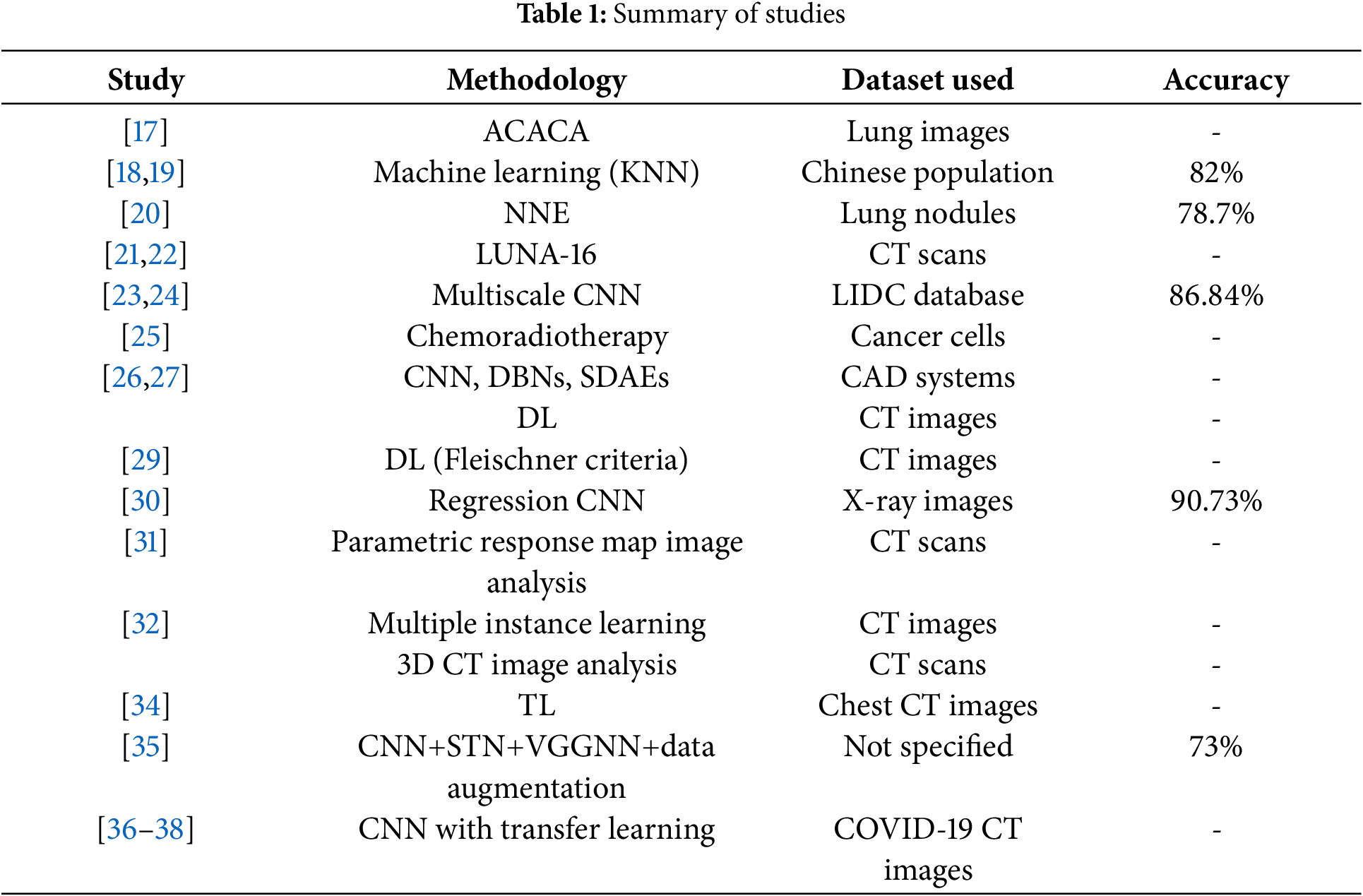
This table encapsulates the diverse approaches and their effectiveness, providing a clear overview of how different models and methodologies compare in the field of lung disease diagnosis through imaging. The literature comprises seven general keywords in the studied papers: sorts of lung sicknesses, DL approach, augmentation of raw data, parameters, and image types. Different specialists could utilize this literature to design their research activities and contributions. The proposed work could additionally enhance the effectiveness, efficiency and increment the DL-oriented detection schemes.
DL models acquire the ability to perform a variety of tasks, including image classification, text analysis, and sound analysis [39]. DL models can rapidly achieve high accuracy, frequently outperforming human performance. To achieve high accuracy and superior results, training a DL neural network is a time-consuming process. DL is capable of performing pattern recognition tasks efficiently; however, the performance of deep networks is difficult to explain.
The model depicted in Fig. 1 is used to detect CB. This method classifies lungs in four distinct steps. The first step is to acquire images; following that, the augmentation technique is used to increase the number of data samples available to feed a neural network. By generating new data samples, data augmentation assists the network in overcoming overfitting. The image is processed by neural networks that learn the features of the image and store them for prediction. Following image processing, classification of the acquired features is performed, and at the conclusion of classification, diagnoses are made.
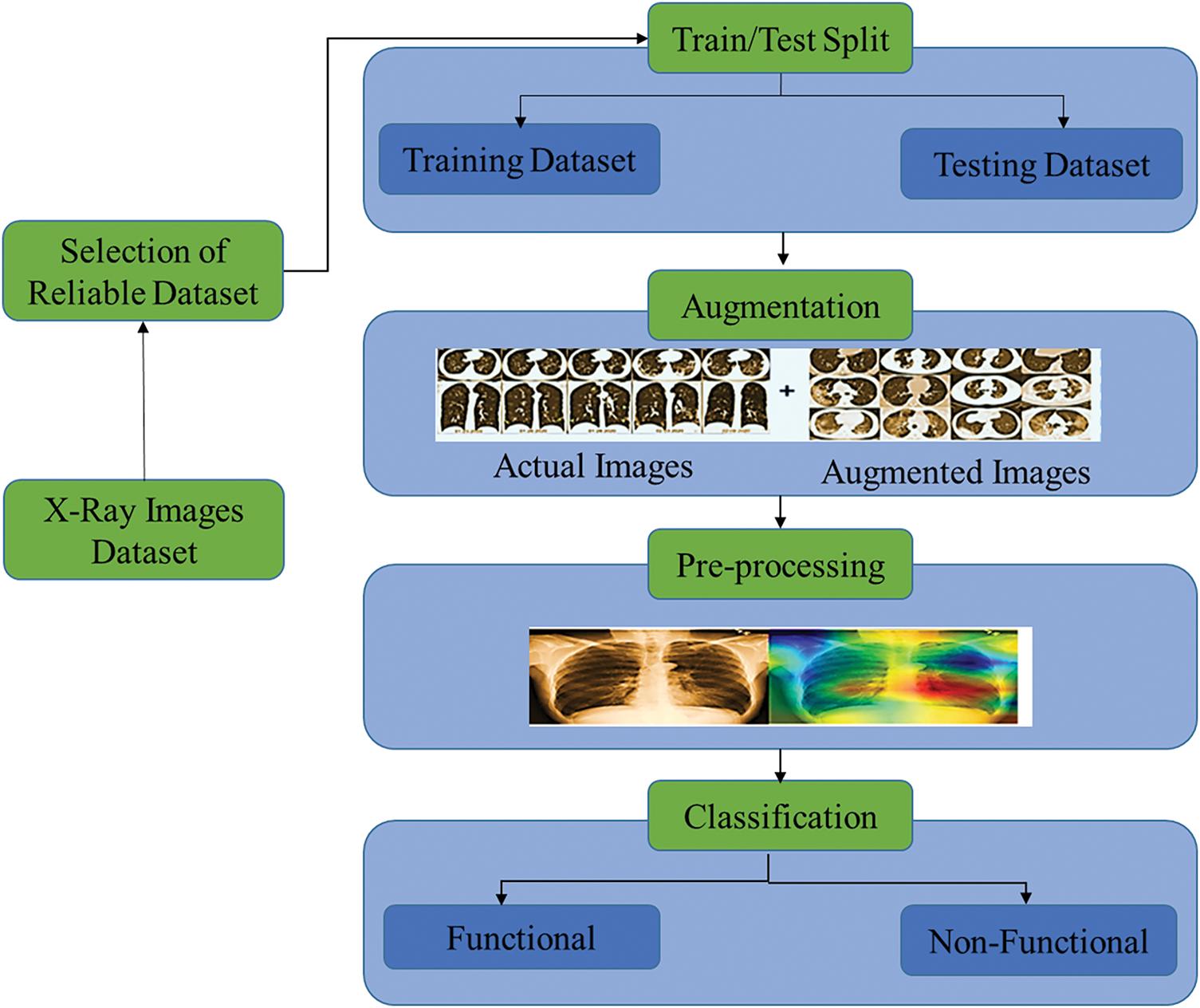
Figure 1: Proposed model for detection of chronic bronchitis
The dataset is required in large quantities and is extremely difficult to collect, primarily for medical diagnosis tasks. To address this issue, we used the data augmentation technique to increase the quantity of data by adding slightly modified copies of existing data and newly generated synthetic data from the given data [40]. The data augmentation technique aids the model in reducing the rate of overfitting and also helps with model regularization during training. The technique of data augmentation is analogous to oversampling.
3.1 Convolutional Neural Network
Neural network-based learning has piqued researchers’ interest in recent years due to its impressive performance in image/video and speech recognition applications. DL’s success is due to powerful computing hardware and a massive amount of labeled training data. Performance can be improved by adding additional layers. Deep networks are used to describe the features extracted from neural networks.
Recent advancements in computing power and modern DL techniques have resulted in the development of a new generation of DL techniques known as CNNs, which have been successfully applied to image recognition, image segmentation, image classification, object detection, and medical diagnosis [41]. CNN is represented in a variety of ways, including by AlexNet, Visual Geometry Group (VGG), GoogleNet, Inception, and ResNet [42]. It is a more refined version of the feed-forward neural network, which is also known as the multi-layer perceptron. CNN has a learning architecture that may be used for imaging diagnostics [43]. To solve these types of problems, a specialized set of knowledge is required, and the stages vary according to the problem. Automated feature detection and extraction can be carried out with high execution results. A few effective formulas for feature extraction are provided below:
CNN contains a layered structure; the input layer is the first layer that takes an image as input and passes it to the next convolution layer. As the image passes through the convolutional layer, it becomes abstracted to the feature map. The convolutional layer contains pooling layers to pick up the essential features from input and reduce the dimensionality of data by merging the outputs at one layer to the next layer’s neurons. The pooling layer uses filters called convolutions filters. Convolutions filters are used in every dataset to extract feature maps n, also called channels. Extracted features are stacked in the matrix.
The hidden layer of CNN is composed of multiple pooling and convolutional layers. Rectified linear unit (ReLU) runs on neuron which is non-linear activation function also known as ramp function [25] (see Eqs. (1) and (2)). The convolution layer is composed of filters that capture input images’ features, and d is the dimension of n feature vectors. Features are extracted, and a vector is generated which contains the features. Eq. (3) demonstrates the feature vector.
Max-pooling operation eliminates nonmaximal values and reduces the computation of the next layers. Features obtained from the max-pooling operation are fed to a fully connected layer.
The fully connected layer is the last layer of CNN; fully connected is prone to overfitting due to occupying most parameters. To reduce the overfitting, dropout is used in the convolution layer. At every individual training step, most particular nodes are further dropped out of the net with the probability of 1-p or only retained with probability p; after dropping out, a depressed net is left, fed further to the next layer. Dropping out reduces the overfitting risk, reduces the number of features, and keeps just only those needed for the task. ReLU is non-saturating activation function. The function of this activation function is to map the negative values to zero. Nonlinear properties of decision functions can be increased using ReLU. The ReLU activation function formula is as follows (see Eq. (5)):
Here x is defined as a result of weighted priority multiplication and paranoid addition, and y is an output of the activation function. The derivative of the ReLU is 0 if x is less than zero; otherwise, it is 1. ReLU can quickly eliminate the problem of Gradient sigmoid activation.
The fully connected layer comes after several convolutional and max-pooling layers. Neurons in a fully connected layer have a connection to previous activation functions. To learn features from convolutions fully connected layer is responsible for learning features. The last layer in the CNN is the classification layer [43]. The SoftMax activation function is used at the classification layer, which brings the values between 0 and 1; loss is calculated after the SoftMax activation function. The architecture of CNN is given in Fig. 2.
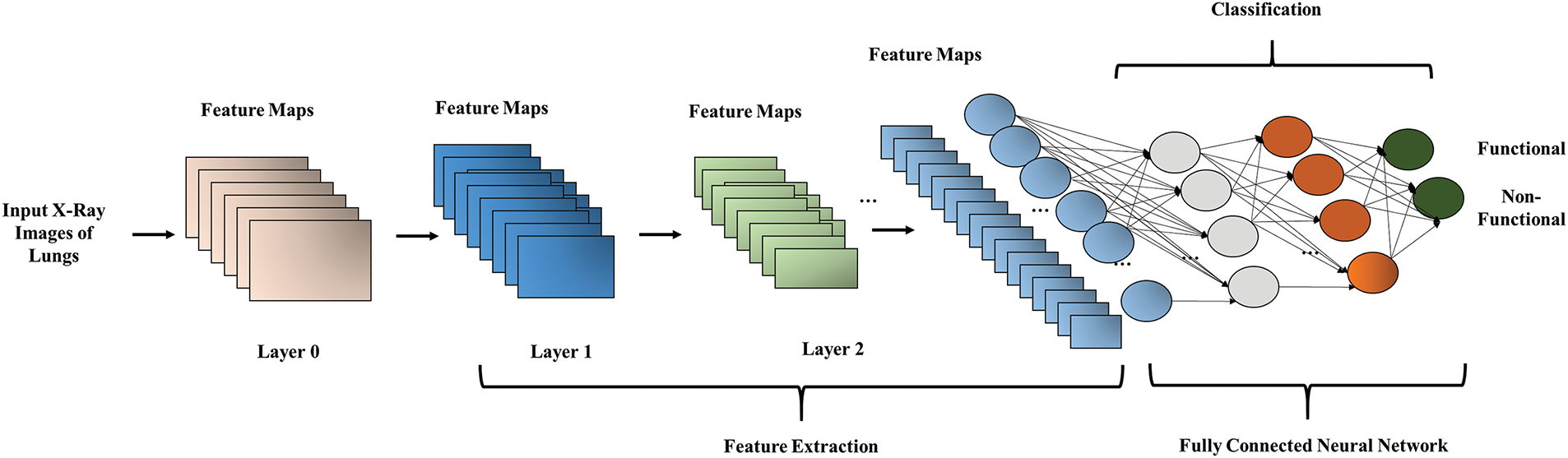
Figure 2: Overview of convolutional neural network architecture for classification
The input of the CNN is composed of four canonical views of X-ray images. X-ray images are analyzed with CNN composed of three neural networks followed by max-pooling functions; every phase size of the image is reduced. At the last of convolutional layers, two fully connected networks, the first network is composed of 1024 neurons and the size of the second variable size depends on the defined problem: classification, multiclass classification, or regression. The loss function can be seen below (see Eq. (6)).
3.1.1 Loss Function of Neural Network
C is the cost function in the above equation, w is the weight, and b is biased, n is the number of samples in the dataset, x is the pixels of images taken as the input parameter, and a is the output value. To update the weights of the neural network backpropagation is applied, backpropagation minimizes the difference between the predicted and real value of the model and tries to increase the accuracy of the model. To prevent the models from overfitting sum of all weights is divided by 2n. Dropping out is also used to prevent a model from overfitting, which randomly shields neurons before the backpropagation.
Cross entropy is used with the SoftMax activation function; backpropagation is applied to reduce neural network loss. Backpropagation goes forward and calculates the loss, then again iterates backward to reduce the loss and it keeps going to minimize the loss and maximize the results. Eq. (7) of backpropagation is below:
In above equation,
Detection of CB in patients relies mostly on CT scans, but the proposed solution uses X-ray images for detection. To solve the proposed problem, CNN needs to be trained by using input X-ray images with their respective labels. As shown in Fig. 3, CNN is trained by using their labels with yes CB and no CB.
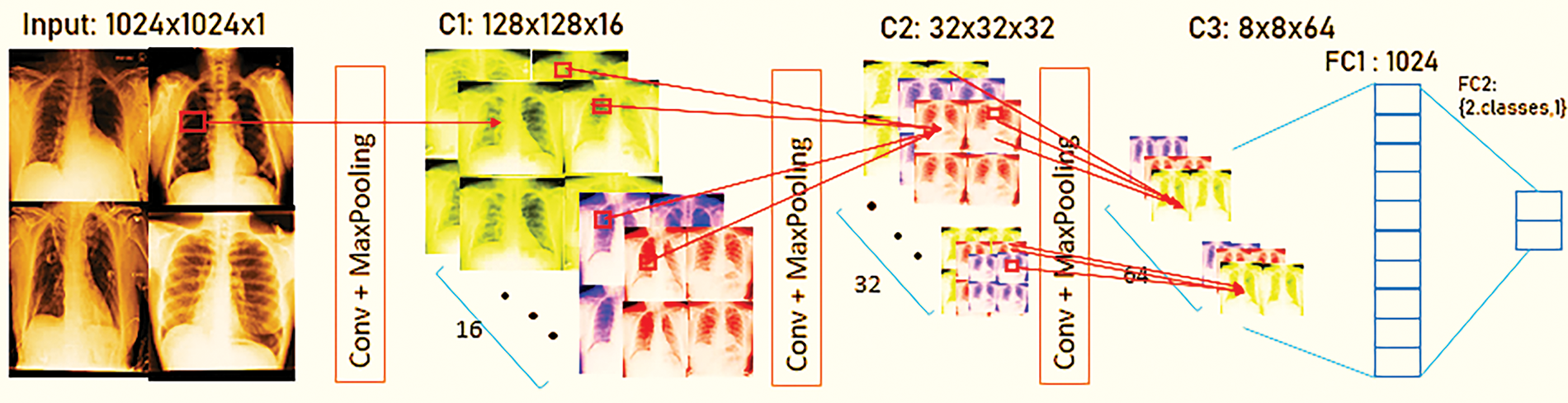
Figure 3: Pre-processing for chronic bronchitis classification
The Algorithm 1 for the proposed work is given below. The first step involves the data preprocessing of input X-ray images. In the second step, features are extracted from input images, and the last step involves the classification of images.

In the proposed approach, features are extracted from X-ray images, and these features are feed to the neural network for the classification of CB.
The experimental analyses were conducted on a specified dataset utilizing a Lenovo Mobile Workstation. This workstation is equipped with a 12th Generation Intel Core i9 processor, operates on Windows 11, and includes 128 GB DDR4 memory, a 2 TB SSD for storage, and an NVIDIA RTX A4000 graphics card. For the computational tasks, the experiments leveraged the Python library TensorFlow, a widely used library in ML to take advantage of its robust capabilities in handling DL models, which are essential for the processing and analysing of the complex datasets used in this research.
The National Institutes of Health (NIH) Chest X-ray Dataset is a significant resource for medical research, featuring 112,120 X-ray images sourced from 30,805 unique patients. Chest X-ray images are among the most common and cost-effective diagnostic tools but pose challenges in clinical diagnosis, often requiring more advanced imaging techniques like CT scans for clarity. The dataset addresses a critical gap in the availability of large, annotated X-ray images’ datasets, which hinders the development of effective computer-aided detection and diagnosis (CAD) systems in real-world medical settings [26].
Due to the extensive number of images, manual labelling is impractical; hence, the dataset utilizes natural language processing (NLP) to derive disease labels from radiological reports, achieving an accuracy of over 90%. The full dataset encompasses 15 class labels for this study, our analysis is focused on a subset, specifically collecting lungs related diseases dataset using classes: pneumonia and emphysema utilized 62 and 127 images respectively, totalling 189 images used in our research. This selective approach allows for a targeted investigation of adducted conditions, contributing to refined diagnostic and prognostic models. The dataset is available at Kaggle. NIH chest X-ray dataset that is publicly available on website is used, and all procedures were conducted in accordance with guidelines that ensure patients’ anonymity and data confidentiality.
X-ray images draw pictures of the inside of the body. White and black shades show different parts of the body. X-ray imaging examination is considered a valuable medical tool for examinations. The most common use of X-ray images is checking the fracture. However, X-ray images can also be used in other diagnoses to detect different lung diseases [44].
3.4.2 Training and Testing Split
The X-ray images dataset is divided into two no overlapping samples to be used for training and validation, and then testing of the model. The training set is fed to the neural network for training; it took time to learn from training data. Validation data is used for fine-tuning the hyperparameters of the model. To prevent a model from overfitting validation set is used. Also, it is used to compare the performances and decide which parameters should be taken further. Testing data is used to evaluate the model after the training and validation phases; testing data are picked, which are not used to optimize the model. Testing data is used to measure the performance of the model. In the training phase, 80% of the data is utilized for training and validation and then 20% of the data is used in the testing phase.
The NIH dataset available on Kaggle only includes 189 X-ray images, which were not sufficient to detect the adduced disease. To overcome this limitation, we employed a data augmentation technique. The proposed approach applies data augmentation to the training dataset after the training/testing split to achieve two primary goals: balancing the classes in the dataset and enhancing the dataset for better modeling. The data augmentation process is explained in Algorithm 1. Ensuring an abundance of data is vital for high-quality results, enabling the productive deployment of most DL models. Data augmentation is a method to artificially enhance data from existing data. Fig. 4 shows the original image vs. images generated using the data augmentation technique. An augmented image retains the same features and intensity as the original image.
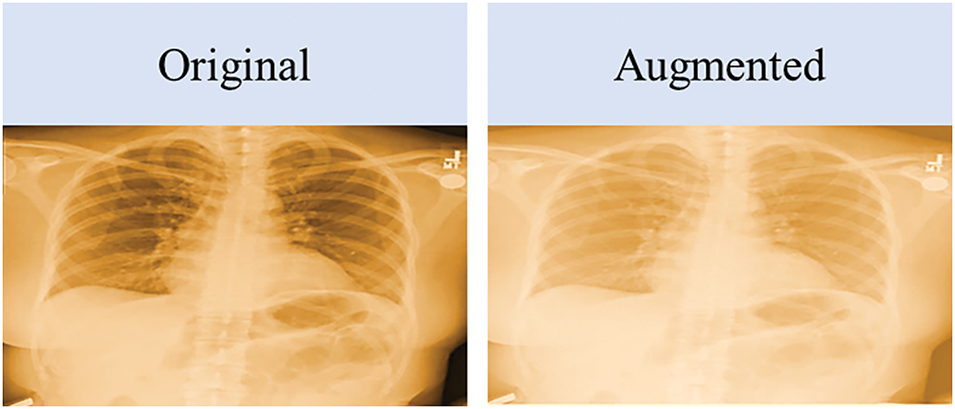
Figure 4: Original image vs. augmented image
Data augmentation is a critical technique used to enhance the volume and diversity of training data for DL models, particularly when original datasets are insufficient to train these models effectively. In our study, data augmentation serves two essential purposes:
• Balancing the Dataset: Addressing class imbalance by generating additional samples for the minority class, ensuring both classes are equally represented in the training phase.
• Enhancing the Dataset: Creating a more diverse dataset to enable the model to generalize better across unseen data, thus improving its robustness and accuracy.
To address these purposes, we employed a variety of augmentation techniques designed to simulate different scenarios that a model might encounter in real-world applications. In this study, a variety of augmentation techniques were employed to enhance the dataset and improve model performance. These geometric transformations include noise injection, which introduces Gaussian noise at varying intensities (low, medium, high, and very high) to simulate real-world variations; contrast adjustment, modifying image contrast by decreasing or increasing it by 20%, 40%, and 60%; and brightness adjustment, which alters image brightness similarly to improve robustness. Scaling and zooming were applied to scale images by factors such as 0.8×, 1.2×, 1.5×, and 2.0×, while elastic deformation was used to mimic non-rigid distortions with varying intensity levels. Additional techniques include Gaussian blur, sharpening, edge detection, embossing, and geometric transformations like skewing (left, right, forward, backward) and shearing along the X- and Y-axes by 15 degrees. These techniques collectively expand the diversity of the dataset, ensuring better generalization and improved model robustness. Fig. 5 illustrates the images created using data augmentation techniques.
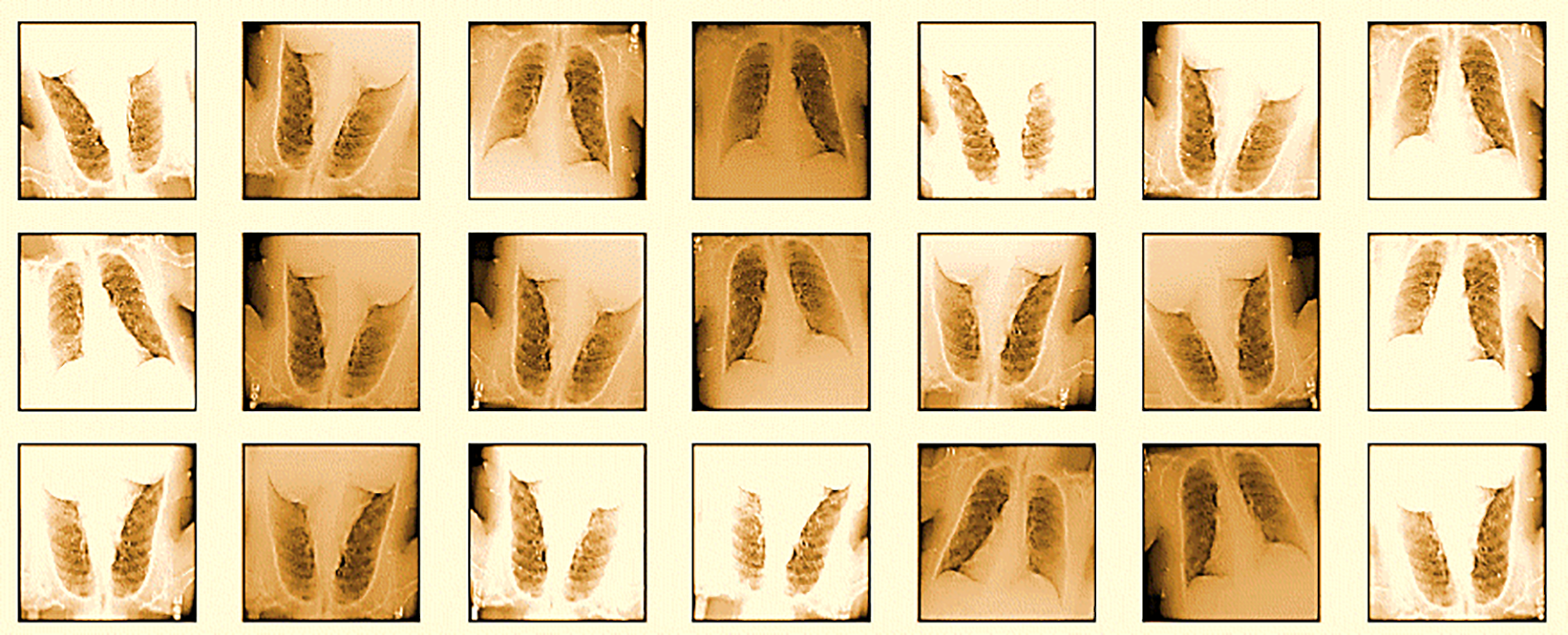
Figure 5: Images created by data augmentation techniques
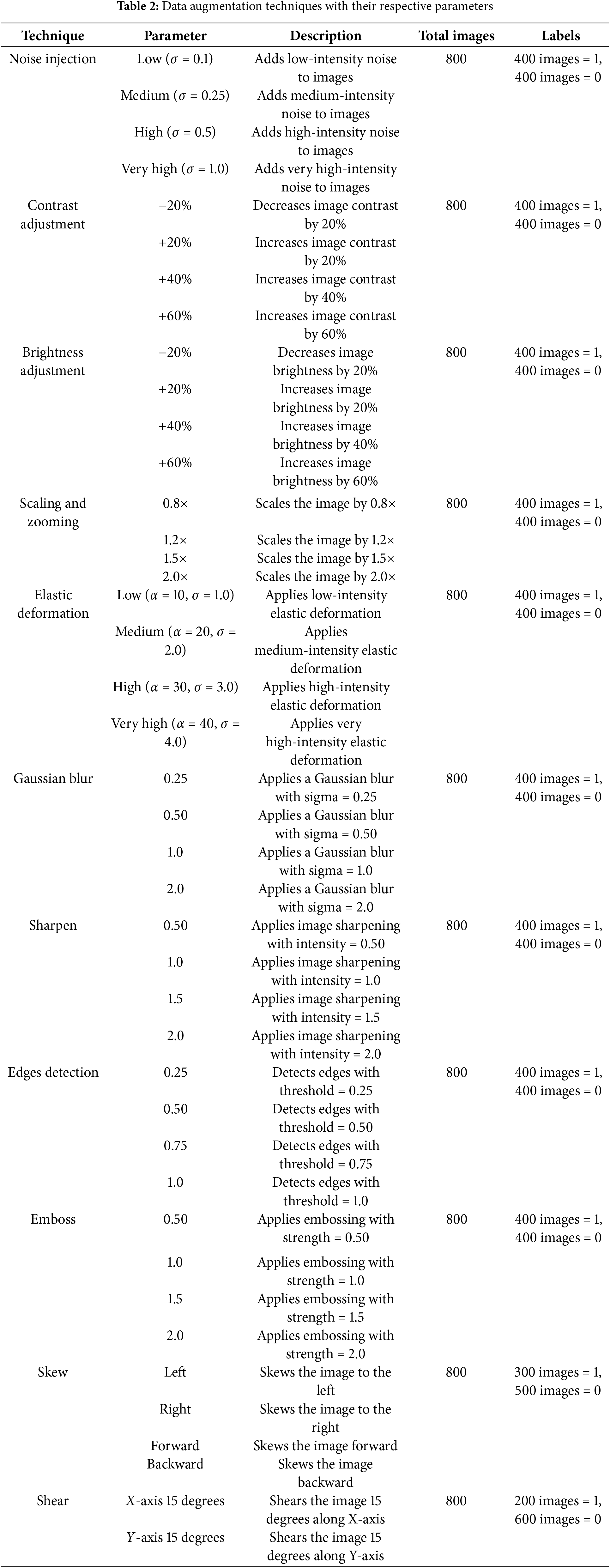
Table 2 provides a comprehensive overview of each data augmentation technique, including the specific parameters used and the distribution of labels across the augmented images.
3.4.4 Segmentation and Feature Extraction
The noise removal process is also very crucial for improving the clarity and quality of the images. This study pre-processes images by filtering out random noise and enhancing image features. This pre-processing step helps reduce the likelihood of overfitting and improves the model’s performance by focusing on relevant features within the images. Techniques such as Gaussian blur and image sharpening are particularly effective in smoothing out noise while preserving essential details, making the images more suitable for accurate analysis by the DL models. Additionally, image segmentation techniques are employed to isolate and analyze specific regions of interest within the images, further enhancing the effectiveness of feature extraction. This step is vital as it allows for the precise extraction of key features from segmented areas, improving the accuracy of the subsequent analysis by the DL models. Fig. 6 depicts the proposed work’s flowchart.
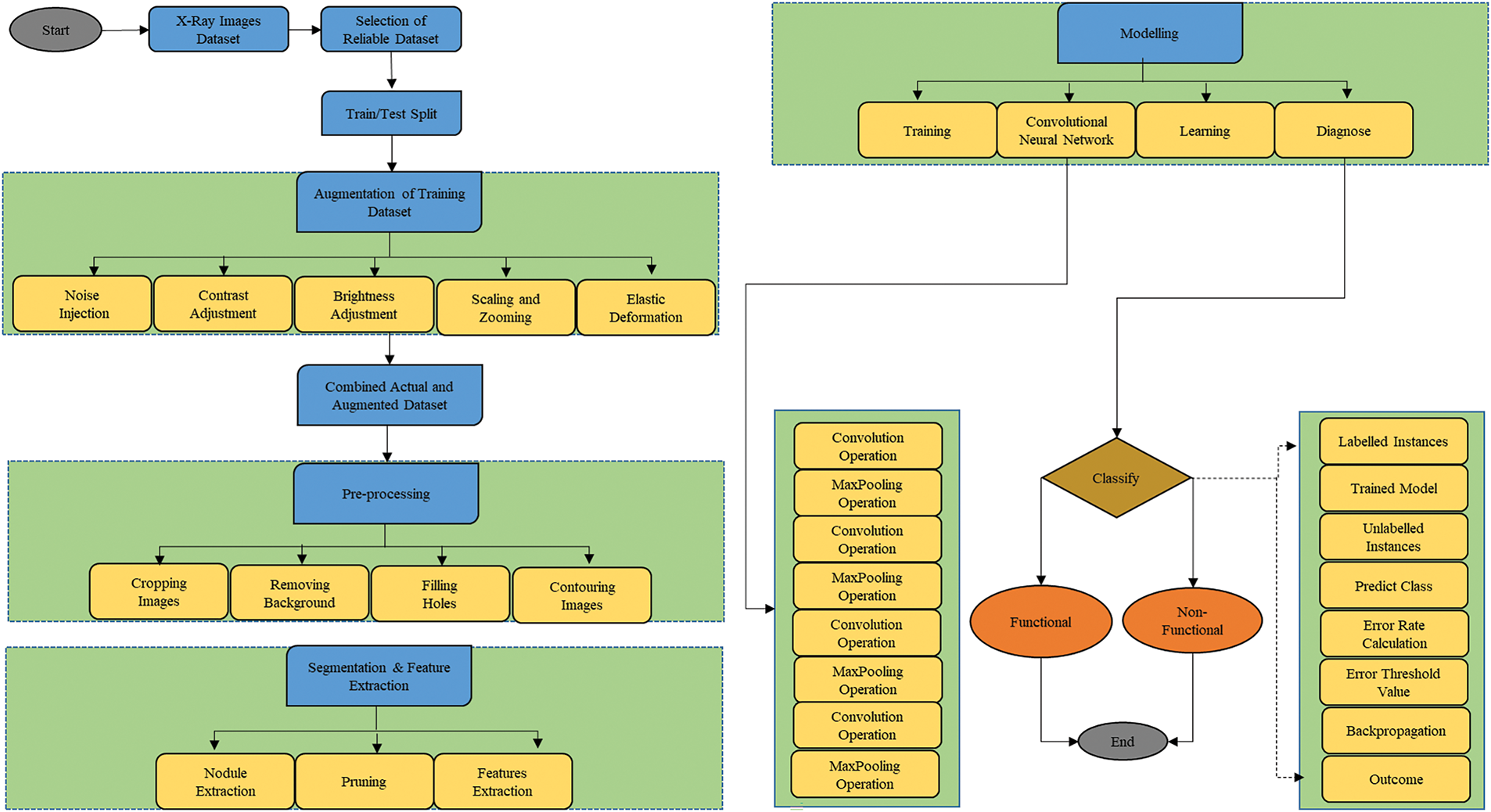
Figure 6: Flowchart for proposed work
3.4.5 Construction of Convolutional Neural Network
To optimize the feature extraction from the input image, we have carefully set the batch size of the neural network to 16 to balance computational load and training efficiency. The kernel size is configured at 3 × 3 to effectively capture fine details and spatial hierarchies in the image data. Each convolution layer is enhanced with 32 filters and followed by a 2 × 2 pooling layer to reduce dimensionality while retaining essential information, allowing the network to delve deeper without significant information loss. Post convolution, a flattened layer compresses the multi-dimensional input into a single dimension, preparing it for dense layer processing. Feature scaling in the fully connected layer is managed through the ReLU activation function to introduce non-linearity and aid in complex pattern learning. Given the binary nature of our classification task, the sigmoid activation function is employed to output probabilities, which are crucial for the final decision-making process. The entire model is refined using the Adam optimizer for its efficient adaptive learning rate adjustments, and loss is computed via binary cross-entropy to address the challenges of binary classification. This architecture and parameter selection are justified by their proven efficacy in enhancing model performance and preventing underfitting, especially when limited data is available. Fig. 7 provides the view of CB classification.
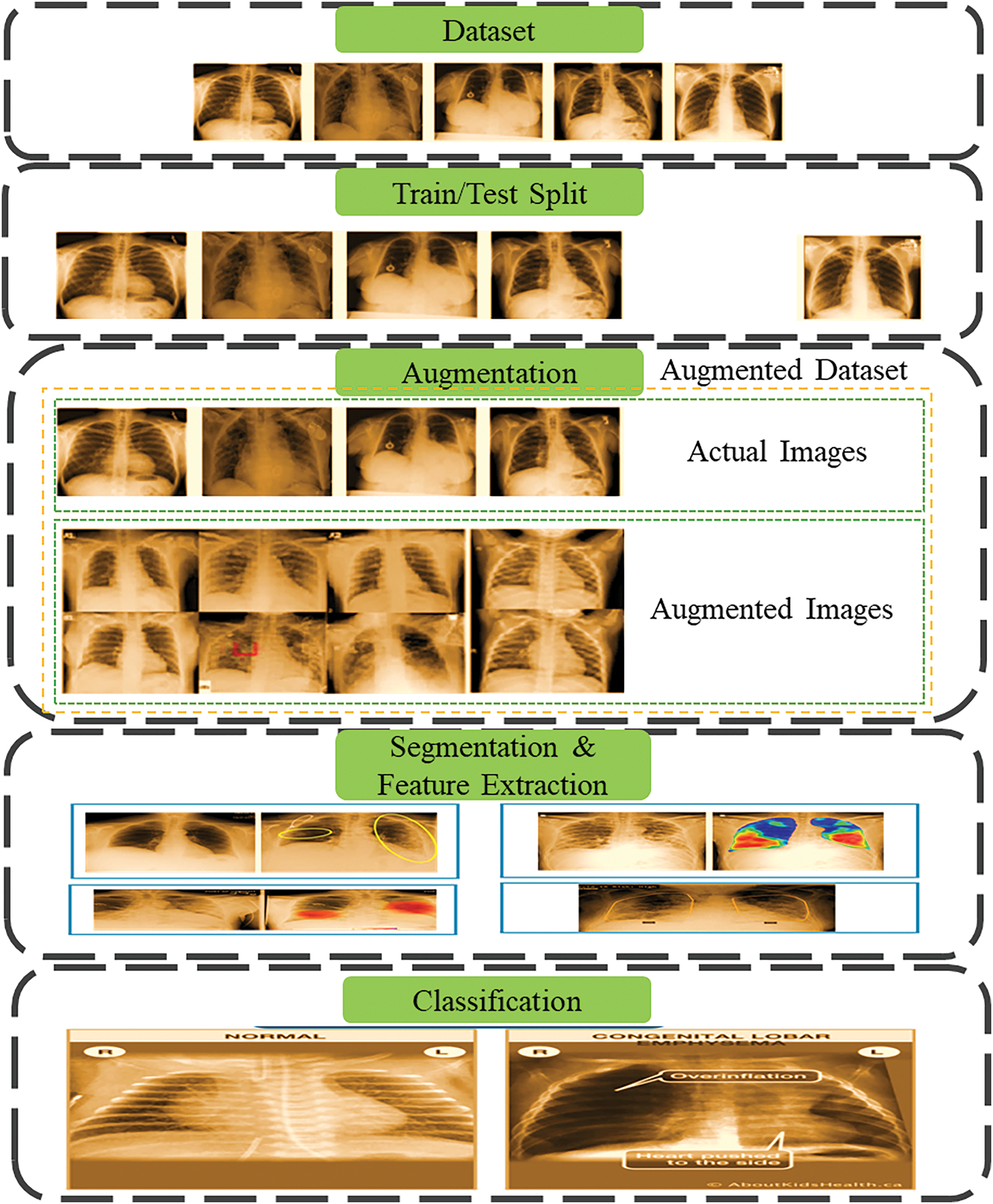
Figure 7: Phases of chronic bronchitis classification
The proposed model’s performance can be quantified in terms of accuracy, validation accuracy, loss, and validation loss. The accuracy of the model can be defined as the percentage of correctly predicted images over the test dataset (see Eq. (8)). The accuracy of validation is used to determine the skill of a trained model in order to fine-tune model parameters. Loss specifies the model’s penalties. Loss quantifies how well a model predicts a single sample. Although validation loss is identical to training loss, it is not used to update the model’s parameters.
TP and TN are the True Positive and True Negative. TP is the number of true samples identified by the model. TN is the number of correctly classified samples.
Using the above loss function (see Eq. (9)) inputs xi and comparing the outputs of your model ŷi with the ground truth values yi using this loss function. This section explains the results obtained from the proposed system.
Our proposed DL model, employing CNNs, achieves an exceptional model accuracy and validation accuracy of 99.2% after training over 25 epochs shown in Fig. 8. The training process reveals a rapid ascent in accuracy within the initial 5 epochs due to the substantial number of samples per epoch, quickly reaching near-perfect levels. Following this initial rise, both the training and validation accuracies stabilize, indicating that the model has effectively learned from the training data without significant overfitting. This plateau in performance demonstrates the model’s robust generalization capabilities, maintaining a high validation accuracy across the span of training, thereby underscoring its effectiveness in handling unseen data. The detailed statistics are shown in Table 3 below.
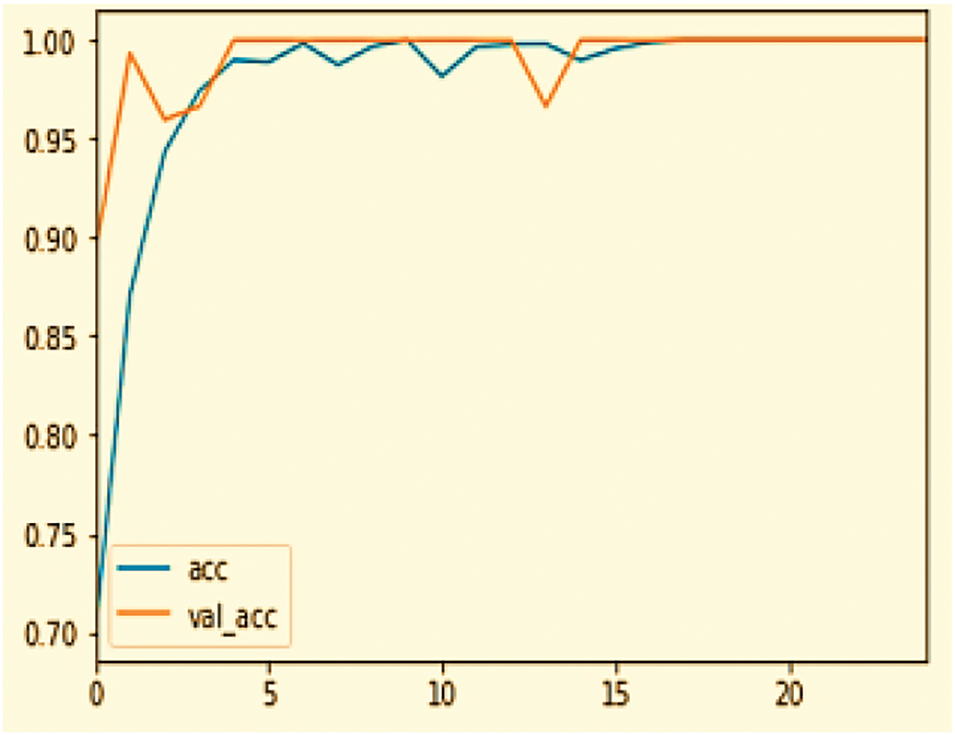
Figure 8: Accuracy curve
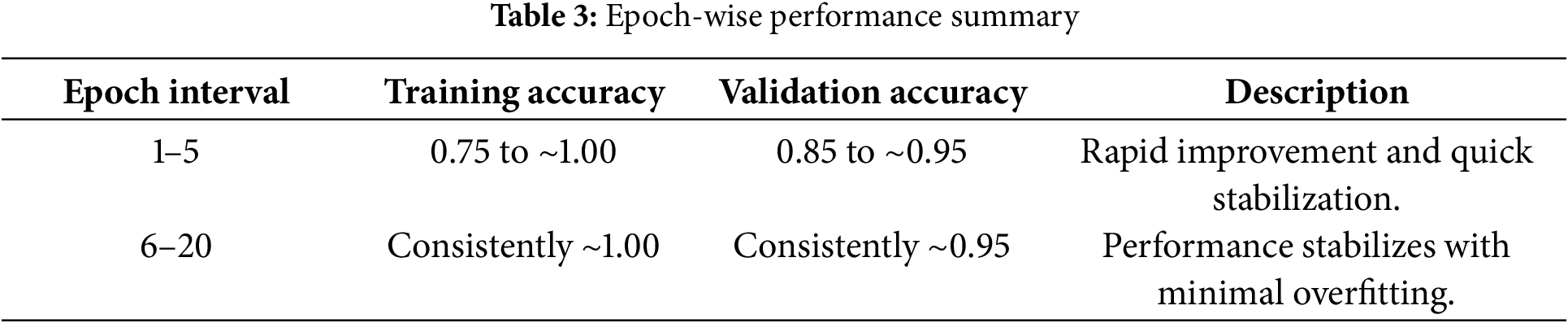
This table outlines the model’s accuracy performance across different stages of the training, highlighting the quick adaptation to the training data and stable generalization to new data.
In the analysis of a DL model’s performance, as shown in the Fig. 9 graph over 25 epochs, both the training loss and the validation loss decrease significantly at the outset, indicating effective error reduction on the dataset. The training loss stabilizes close to zero within the initial epochs, demonstrating that the model has thoroughly learned the training data. However, the validation loss, despite its overall downward trend, exhibits slight fluctuations, particularly around the 10th and 15th epochs, reflecting some inconsistency in the model’s performance on new data. This pattern suggests that the model generalizes well overall, balance between low loss for accuracy and avoiding too little loss to prevent overfitting is crucial for optimizing model performance. The detail statistics are shown in Table 4 below.
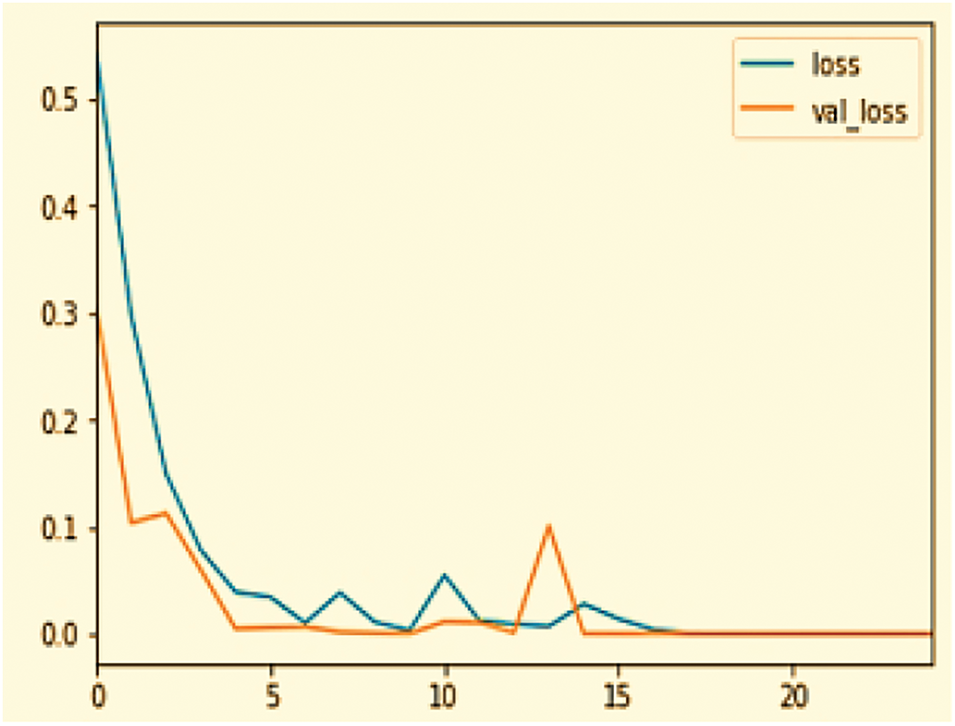
Figure 9: Loss curve
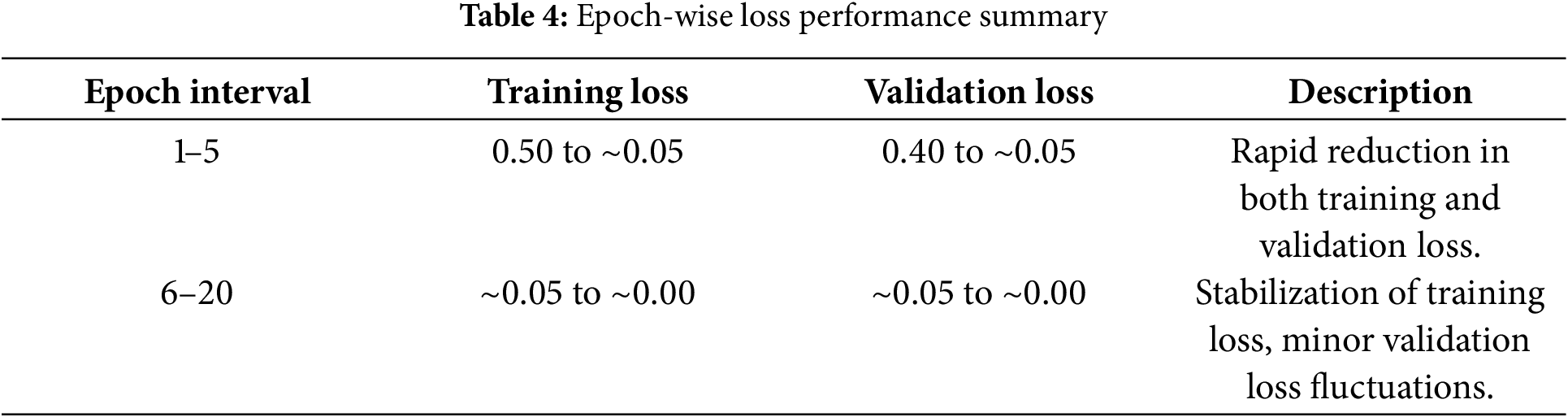
This table summarizes the loss performance, highlighting the initial rapid learning phase and subsequent stabilization, indicating effective model training and generalization.
The accuracy and loss curves over 20 epochs reveal key insights into model performance. Training accuracy quickly stabilizes near 1.00, while validation accuracy initially dips, suggesting slight overfitting, but recovers and aligns closely with training accuracy, indicating effective generalization adjustments. Loss metrics decrease sharply, with training loss nearing zero quickly. However, validation loss shows fluctuations, notably a significant spike around the 15th epoch, hinting at potential overfitting issues which are subsequently addressed as the loss decreases again. These patterns suggest that while the model learns effectively, there is room for improvement in managing overfitting, possibly through techniques like learning rate adjustments or regularization.
The augmentation techniques played a critical role in mitigating these challenges by increasing dataset diversity and reducing the risk of overfitting. Noise injection, contrast and brightness adjustments, and elastic deformation helped the model become more robust to variations, enhancing its ability to generalize across unseen data. Techniques like scaling and zooming ensured the model could handle different image resolutions while skewing and shearing improved its adaptability to geometric distortions. As a result, these augmentations contributed to the eventual stabilization of validation accuracy and the decline in validation loss, showcasing their effectiveness in improving the model’s performance and robustness. Nevertheless, the observed fluctuations highlight opportunities for further refinements, such as fine-tuning learning rates or introducing additional regularization techniques.
Table 5 summarizes the other performance metrices, highlighting the effective model training and validation. The progression of the model is assessed using performance metrices including precision, recall, and F1-score, by using the following formulas (Eqs. (10) to (12)).
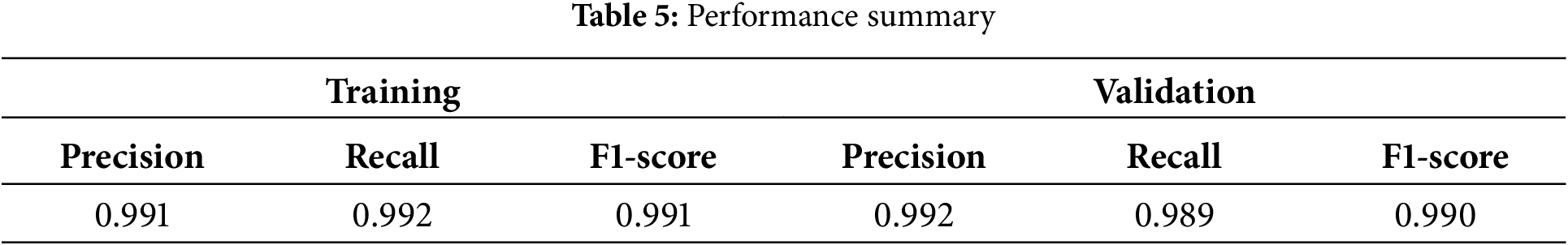
This research provides insights into the CNN model, which we used to detect CB in healthy individuals and those with a functional state of CB by analyzing the lung’s X-ray images. It is self-evident that data is critical in any DL framework; if the data is more precise and specific about CB indications, it will generate the highest precision with the best results. The experimental results demonstrate that the proposed work is extremely precise. As a result, if it is used in real-time situations, it will be extremely helpful in diagnosing CB. The results indicate that the proposed approach is capable of assisting in the timely and accurate detection of CB in addition to recognizing its precise position. Following that, the proposed method is more appropriate for CB detection from X-ray images. The test results achieved a precision of 99.2%, demonstrating the proposed work’s suitability for discriminating between functional and non-functional states of CB. As a result, the proposed strategy for incorporating a clinical expert system (for primary diagnosis and screening) by radiologists or clinical specialists is reasonable. Table 6 compares the proposed approach to existing approaches.
• The p-values obtained from paired t-tests compare the accuracy of each previous method with the proposed CNN model.
• A p-value less than 0.05 indicates a statistically significant improvement in the performance of the proposed model over the respective previous method.
6 Conclusion, Limitations and Future Work
Lungs are essential organs responsible for transporting oxygen throughout the body and eliminating harmful substances. Various factors can damage the lungs, leading to severe diseases that cause numerous deaths annually. Early detection and treatment are crucial in preventing fatalities. This article presents a method for detecting CB using X-ray images. A CNN achieved a 99.2% accuracy in identifying CB through these images. The technique leverages data augmentation to enrich the dataset, enhancing the CNN’s predictive accuracy with each training epoch. This study not only aids physicians in diagnosing CB at early stages, potentially saving lives, but also sets the groundwork for future improvements. Enhancements could include refining the classifier’s methodology, integrating multiple classifiers, and implementing more efficient feature extraction techniques.
The model was initially trained and validated on a restricted set of image variations, which limits its application to a broader range of clinical scenarios. Specifically, this study utilized only a small subset of the NIH dataset, which may not fully represent the variability encountered in clinical practice. To enhance the model’s generalizability and reliability in real-world conditions, future research should encompass a more extensive collection of both pathological and non-pathological images from diverse demographics and clinical environments. Additionally, integrating multimodal data such as combining radiographic imaging with electronic health records and genomic data could significantly enrich the diagnostic capabilities of the tool. Plans to test the enhanced model on independent datasets will also be crucial for validating its efficacy across different clinical settings. Exploring the use of Quantum Machine Learning (QML) for synthesizing datasets could also be transformative. QML offers the potential to process data in higher dimensions and at greater speeds, thereby improving the efficiency and depth of data analysis for healthcare applications.
Acknowledgement: We extend our gratitude to our institutions for their moral support.
Funding Statement: This study did not receive any funding.
Author Contributions: Conceptualization: Fahad Ahmad, Saad Awadh Alanazi, Kashaf Junaid, Maryam Shabbir; Writing Original Draft: Kashaf Junaid, Maryam Shabbir, Fahad Ahmad, Asim Ali; Software: Fahad Ahmad, Maryam Shabbir, Asim Ali; Investigation: Fahad Ahmad, Saad Awadh Alanazi, Kashaf Junaid; Validation: Fahad Ahmad, Maryam Shabbir, Asim Ali; Visualization: Kashaf Junaid, Fahad Ahmad. All authors reviewed the results and approved the final version of the manuscript.
Availability of Data and Materials: The dataset used in this study is publicly available.
Ethics Approval: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest to report regarding the present study.
References
1. Arakawa H, Niimi H, Kurihara Y, Nakajima Y, Webb WR. Expiratory high-resolution CT: diagnostic value in diffuse lung diseases. Am J Roentgenol. 2000;175(6):1537–43. doi:10.2214/ajr.175.6.1751537. [Google Scholar] [PubMed] [CrossRef]
2. Nicod LP. Lung defences: an overview. European Respir Rev. 2005;14(95):45–50. doi:10.1183/09059180.05.00009501. [Google Scholar] [CrossRef]
3. Thorley AJ, Tetley TD. Pulmonary epithelium, cigarette smoke, and chronic obstructive pulmonary disease. Int J Chronic Obstr Pulm Dis. 2007;2(4):409–28. [Google Scholar]
4. Team NLSTR. Reduced lung-cancer mortality with low-dose computed tomographic screening. New Engl J Med. 2011;365(5):395–409. doi:10.1056/NEJMoa1102873. [Google Scholar] [PubMed] [CrossRef]
5. Agustí A, Edwards LD, Rennard SI, MacNee W, Tal-Singer R, Miller BE, et al. Persistent systemic inflammation is associated with poor clinical outcomes in COPD: a novel phenotype. PLoS One. 2012;7(5):e37483. doi:10.1371/journal.pone.0037483. [Google Scholar] [PubMed] [CrossRef]
6. Rubin LJ, Badesch DB, Barst RJ, Galie N, Black CM, Keogh A, et al. Bosentan therapy for pulmonary arterial hypertension. New Engl J Med. 2002;346(12):896–903. doi:10.1056/NEJMoa012212. [Google Scholar] [PubMed] [CrossRef]
7. Thun MJ, Hannan LM, Adams-Campbell LL, Boffetta P, Buring JE, Feskanich D, et al. Lung cancer occurrence in never-smokers: an analysis of 13 cohorts and 22 cancer registry studies. PLoS Med. 2008;5(9):e185. doi:10.1371/journal.pmed.0050185. [Google Scholar] [PubMed] [CrossRef]
8. Berry CE, Wise RA. Mortality in COPD: causes, risk factors, and prevention. COPD: J Chronic Obstr Pulm Dis. 2010;7(5):375–82. doi:10.3109/15412555.2010.510160. [Google Scholar] [PubMed] [CrossRef]
9. Halpin DM. Mortality of patients with COPD. Expert Rev Respir Med. 2024;18(6):381–95. doi:10.1080/17476348.2024.2375416. [Google Scholar] [PubMed] [CrossRef]
10. Stacher E, Graham BB, Hunt JM, Gandjeva A, Groshong SD, McLaughlin VV, et al. Modern age pathology of pulmonary arterial hypertension. Am J Respi Crit Care Med. 2012;186(3):261–72. doi:10.1164/rccm.201201-0164OC. [Google Scholar] [PubMed] [CrossRef]
11. Garcia-Aymerich J, Gómez FP, Benet M, Farrero E, Basagana X, Gayete A, et al. Identification and prospective validation of clinically relevant chronic obstructive pulmonary disease (COPD) subtypes. Thorax. 2011;66(5):430–7. doi:10.1136/thx.2010.154484. [Google Scholar] [PubMed] [CrossRef]
12. Liu L, Li S, Wu Y, Wang X, Huang D, Pei C, et al. Effect of myrtol on chronic bronchitis or chronic obstructive pulmonary disease: a protocol for systematic review and meta-analysis. Medicine. 2020;99(28):e20692. doi:10.1097/MD.0000000000020692. [Google Scholar] [PubMed] [CrossRef]
13. Adeloye D, Song P, Zhu Y, Campbell H, Sheikh A, Rudan I. Global, regional, and national prevalence of, and risk factors for, chronic obstructive pulmonary disease (COPD) in 2019: a systematic review and modelling analysis. Lancet Respir Med. 2022;10(5):447–58. doi:10.1016/S2213-2600(21)00511-7. [Google Scholar] [PubMed] [CrossRef]
14. Snider GL, Kleinerman J, Thurlbeck WM, Bengali ZH. The definition of emphysema: report of a national heart, lung, and blood institute, division of lung diseases workshop. Am Rev Respir Dis. 1985 Jul;132(1):182–5. doi:10.1164/arrd.1985.132.1.182. [Google Scholar] [CrossRef]
15. Hoffman EA, Simon BA, McLennan G. State of the art. A structural and functional assessment of the lung via multidetector-row computed tomography: phenotyping chronic obstructive pulmonary disease. Proc Am Thorac Soc. 2006;3(6):519–32. doi:10.1513/pats.200603-086MS. [Google Scholar] [PubMed] [CrossRef]
16. Al-qaness MA, Zhu J, AL-Alimi D, Dahou A, Alsamhi SH, Abd Elaziz M, et al. Chest X-ray images for lung disease detection using deep learning techniques: a comprehensive survey. Arch Comput Methods Eng. 2024, 1–35. [Google Scholar]
17. Asif S, Wenhui Y, ur-Rehman S, ul-ain Q, Amjad K, Yueyang Y, et al. Advancements and prospects of machine learning in medical diagnostics: unveiling the future of diagnostic precision. Arch Comput Methods Eng. 2024. doi:10.1007/s11831-024-10148-w. [Google Scholar] [CrossRef]
18. Mansoor A, Bagci U, Foster B, Xu Z, Papadakis GZ, Folio LR, et al. Segmentation and image analysis of abnormal lungs at CT: current approaches, challenges, and future trends. Radiographics. 2015;35(4):1056–76. doi:10.1148/rg.2015140232. [Google Scholar] [PubMed] [CrossRef]
19. Gulati A. LungAI: a deep learning convolutional neural network for automated detection of COVID-19 from posteroanterior chest X-rays. medRxiv. 2020. doi:10.1101/2020.12.19.20248530. [Google Scholar] [CrossRef]
20. Ma X, Wu Y, Zhang L, Yuan W, Yan L, Fan S, et al. Comparison and development of machine learning tools for the prediction of chronic obstructive pulmonary disease in the Chinese population. J Transl Med. 2020;18(1):1–14. doi:10.1186/s12967-020-02312-0. [Google Scholar] [PubMed] [CrossRef]
21. Tu M, Wang X, Liu H, Jia H, Wang Y, Li J, et al. Precision patient selection for improved detection of circulating genetically abnormal cells in pulmonary nodules. Sci Rep. 2024;14(1):22532. doi:10.1038/s41598-024-73542-1. [Google Scholar] [PubMed] [CrossRef]
22. Chen H, Wu W, Xia H, Du J, Yang M, Ma B, editors. Classification of pulmonary nodules using neural network ensemble. In: Advances in Neural Networks–ISNN 2011: 8th International Symposium on Neural Networks, ISNN 2011; 2011 May 29–Jun 1; Guilin, China: Springer. [Google Scholar]
23. Shen W, Zhou M, Yang F, Yang C, Tian J, editors. Multi-scaleconvolutional neural networks for lung nodule classification. In: Information Processing in Medical Imaging: 24th International Conference, IPMI 2015; 2015 Jun 28–Jul 3; Sabhal Mor Ostaig, Isle of Skye, UK:Springer International Publishing. p. 588–99. [Google Scholar]
24. Zatloukal P, Petruzelka L, Zemanova M, Havel L, Janku F, Judas L, et al. Concurrent versus sequential chemoradiotherapy with cisplatin and vinorelbine in locally advanced non-small cell lung cancer: a randomized study. Lung Cancer. 2004;46(1):87–98. doi:10.1016/j.lungcan.2004.03.004. [Google Scholar] [PubMed] [CrossRef]
25. Liang HY, Zhou H, Li XL, Yin ZH, Guan P, Zhou BS. Chemo-radiotherapy for advanced non-small cell lung cancer: concurrent or sequential? It’s no longer the question: a systematic review. Int J Cancer. 2010;127(3):718–28. doi:10.1002/ijc.25087. [Google Scholar] [PubMed] [CrossRef]
26. Sun W, Zheng B, Qian W. Computer aided lung cancer diagnosis with deep learning algorithms. In: Medical imaging 2016: computer-aided diagnosis. SPIE; 2016 Mar. Vol. 9785. p. 241–48. [Google Scholar]
27. Negahdar M, Beymer D. Lung tissue characterization for emphysema differential diagnosis using deep convolutional neural networks. In: Medical imaging 2016: computer-aided diagnosis. SPIE; 2019 Mar. Vol. 10950. p. 950–55. [Google Scholar]
28. Zhang F, Han H, Li M, Tian T, Zhang G, Yang Z, et al. Revolutionizing diagnosis of pulmonary mycobacterium tuberculosis based on CT: a systematic review of imaging analysis through deep learning. Front Microbiol. 2025;15:1510026. doi:10.3389/fmicb.2024.1510026. [Google Scholar] [PubMed] [CrossRef]
29. Humphries SM, Notary AM, Centeno JP, Strand MJ, Crapo JD, Silverman EK, et al. Deep learning enables automatic classification of emphysema pattern at CT. Radiology. 2020;294(2):434–44. doi:10.1148/radiol.2019191022. [Google Scholar] [PubMed] [CrossRef]
30. Altan G, Kutlu Y, Allahverdi N. Deep learning on computerized analysis of chronic obstructive pulmonary disease. IEEE J Biomed Health Inform. 2019;24(5):1344–50. doi:10.1109/JBHI.2019.2931395. [Google Scholar] [PubMed] [CrossRef]
31. Galbán CJ, Han MK, Boes JL, Chughtai KA, Meyer CR, Johnson TD, et al. Computed tomography-based biomarker provides unique signature for diagnosis of COPD phenotypes and disease progression. Nature Med. 2012;18(11):1711–5. doi:10.1038/nm.2971. [Google Scholar] [PubMed] [CrossRef]
32. Cheplygina V, Sørensen L, Tax DM, Pedersen JH, Loog M, De Bruijne M. Classification of COPD with multiple instance learning. In: 2014 22nd International Conference on pattern recognition; 2014; IEEE. p. 1508–13. [Google Scholar]
33. Wu Y, Xia S, Liang Z, Chen R, Qi S. Artificial intelligence in COPD CT images: identification, staging, and quantitation. Respir Res. 2024;25(1):319. doi:10.1186/s12931-024-02913-z. [Google Scholar] [PubMed] [CrossRef]
34. Sharma A, Rani S, Gupta D. Artificial intelligence-based classification of chest X-ray images into COVID-19 and other infectious diseases. Int J Biomed Imaging. 2020;2020(1):8889023. doi:10.1155/2020/8889023. [Google Scholar] [PubMed] [CrossRef]
35. Bharati S, Podder P, Mondal MRH. Hybrid deep learning for detecting lung diseases from X-ray images. Inform Med Unlocked. 2020;20(7):100391. doi:10.1016/j.imu.2020.100391. [Google Scholar] [PubMed] [CrossRef]
36. Zhu J, Shen B, Abbasi A, Hoshmand-Kochi M, Li H, Duong TQ. Deep transfer learning artificial intelligence accurately stages COVID-19 lung disease severity on portable chest radiographs. PLoS One. 2020;15(7):e0236621. doi:10.1371/journal.pone.0236621. [Google Scholar] [PubMed] [CrossRef]
37. Ahmad F, Almuayqil SN, Humayun M, Naseem S, Khan WA, Junaid K. Prediction of COVID-19 cases using machine learning for effective public health management. Comput Mater Contin. 2020;66(3):2265–82. doi:10.32604/cmc.2021.013067. [Google Scholar] [CrossRef]
38. Junaid K, Qasim S, Yasmeen H, Ejaz H, Alsrhani A, Ullah MI, et al. Potential inhibitory effect of vitamins against COVID-19. Comput Mater Contin. 2020;66(1):707–14. doi:10.32604/cmc.2020.012976. [Google Scholar] [CrossRef]
39. Saturi S, Banda S. Modelling of deep learning enabled lung disease detection and classification on chest X-ray images. Int J Healthc Manag. 2022;10(4):1–12. doi:10.1080/20479700.2022.2102223. [Google Scholar] [CrossRef]
40. Wang R, Chen L-C, Moukheiber L, Seastedt KP, Moukheiber M, Moukheiber D, et al. Enabling chronic obstructive pulmonary disease diagnosis through chest X-rays: a multi-site and multi-modality study. Int J Med Inform. 2023;178(9589):105211. doi:10.1016/j.ijmedinf.2023.105211. [Google Scholar] [PubMed] [CrossRef]
41. Moses DA. Deep learning applied to automatic disease detection using chest X-rays. J Med Imaging Radiat Oncol. 2021;65(5):498–517. doi:10.1111/1754-9485.13273. [Google Scholar] [PubMed] [CrossRef]
42. Zainorzuli SM, Che Abdullah SA, Abidin HZ, Ahmat Ruslan F. Comparison study on convolution neural network (CNN) techniques for image classification. J Electr Electron Syst Res. 2022;20:11–7. doi:10.24191/jeesr.v20i1.002. [Google Scholar] [CrossRef]
43. Banerjee I, Ling Y, Chen MC, Hasan SA, Langlotz CP, Moradzadeh N, et al. Comparative effectiveness of convolutional neural network (CNN) and recurrent neural network (RNN) architectures for radiology text report classification. Artif Intell Med. 2019;97(2):79–88. doi:10.1016/j.artmed.2018.11.004. [Google Scholar] [PubMed] [CrossRef]
44. Yamashita R, Nishio M, Do RKG, Togashi K. Convolutional neural networks: an overview and application in radiology. Insights into Imaging. 2018;9(4):611–29. doi:10.1007/s13244-018-0639-9. [Google Scholar] [PubMed] [CrossRef]
45. Mehrotra R, Agrawal R, Ansari M. Diagnosis of hypercritical chronic pulmonary disorders using dense convolutional network through chest radiography. Multimed Tools Appl. 2022;81(6):7625–49. doi:10.1007/s11042-021-11748-5. [Google Scholar] [PubMed] [CrossRef]
46. Rahman T, Khandakar A, Kadir MA, Islam KR, Islam KF, Mazhar R, et al. Reliable tuberculosis detection using chest X-ray with deep learning, segmentation and visualization. IEEE Access. 2020;8:191586–601. doi:10.1109/ACCESS.2020.3031384. [Google Scholar] [CrossRef]
47. Chen K-C, Yu H-R, Chen W-S, Lin W-C, Lee Y-C, Chen H-H, et al. Diagnosis of common pulmonary diseases in children by X-ray images and deep learning. Sci Rep. 2020;10(1):17374. doi:10.1038/s41598-020-73831-5. [Google Scholar] [PubMed] [CrossRef]
Cite This Article
 Copyright © 2025 The Author(s). Published by Tech Science Press.
Copyright © 2025 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF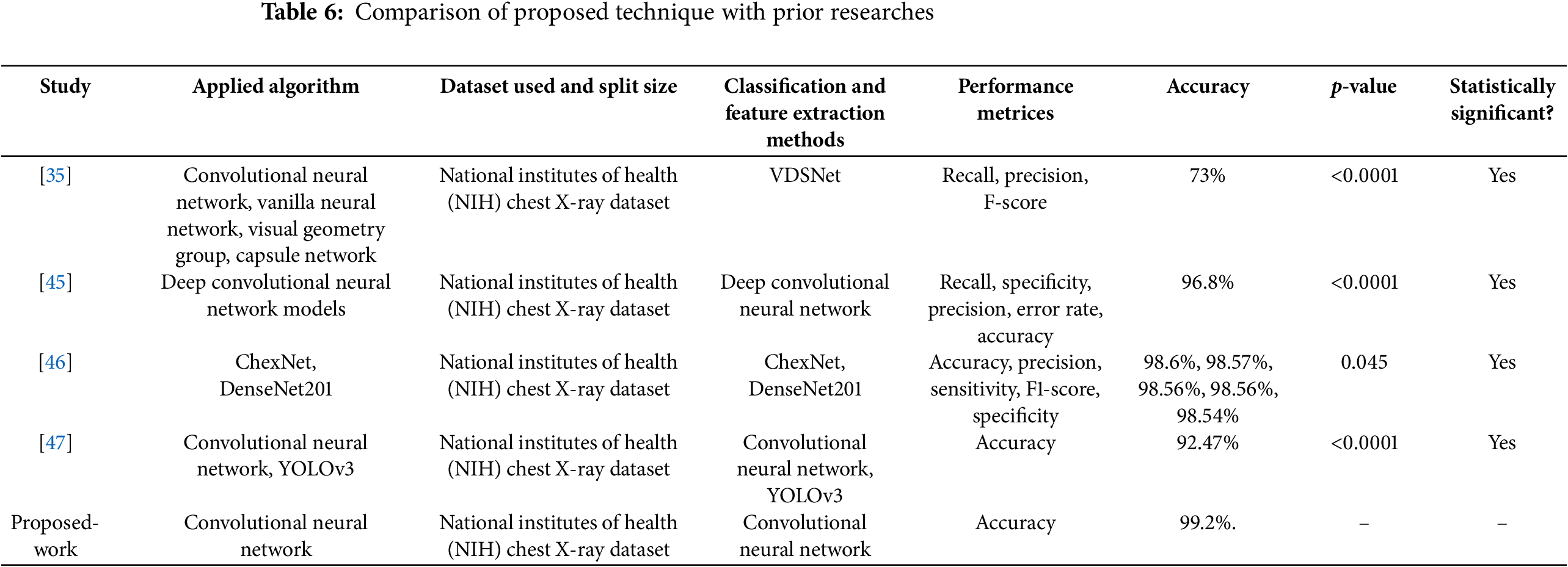
 Downloads
Downloads
 Citation Tools
Citation Tools