 Open Access
Open Access
ARTICLE
Research on Multimodal Brain Tumor Segmentation Algorithm Based on Feature Decoupling and Information Bottleneck Theory
1 China Academy of Engineering Physics, Graduate School, Beijing, 100193, China
2 Shanghai Zhangjiang Institute of Mathematics, Biomedical Laboratory, Shanghai, 201203, China
3 Institute of Applied Physics and Computational Mathematics, Tang Chuang Centre, Beijing, 100094, China
* Corresponding Author: Junping Yin. Email:
(This article belongs to the Special Issue: Novel Methods for Image Classification, Object Detection, and Segmentation)
Computers, Materials & Continua 2025, 82(2), 3281-3307. https://doi.org/10.32604/cmc.2024.057991
Received 02 September 2024; Accepted 06 December 2024; Issue published 17 February 2025
Abstract
Aiming at the problems of information loss and the relationship between features and target tasks in multimodal medical image segmentation, a multimodal medical image segmentation algorithm based on feature decoupling and information bottleneck theory is proposed in this paper. Based on the reversible network, the bottom-up learning method for different modal information is constructed, which enhances the features’ expression ability and the network’s learning ability. The feature fusion module is designed to balance multi-directional information flow. To retain the information relevant to the target task to the maximum extent and suppress the information irrelevant to the target task, the feature decoupling module is designed to ensure a strong correlation between the feature and the target task. A loss function based on information bottleneck theory was intended to improve information quality and remove redundant information. Based on BraTs2021, BraTs2023-MET and ANNLIB datasets, the proposed algorithm is analyzed qualitatively and quantitatively in this paper. In the quantitative experiment, the Dice coefficient of the proposed algorithm was increased by 0.110 on average compared with other methods, and the HD95 was decreased by 28.568 on average compared with other methods. In qualitative analysis, the proposed algorithm can effectively segment the incoherent region between the lesion and the lesion boundary and achieve accurate segmentation of the lesion.Keywords
In recent years, with the development of artificial intelligence algorithms, data-driven algorithms are constantly changing from single-modal data to multimodal data. The theories and methods in mathematics and information provide theoretical support for studying the mechanism of multimodal data fusion and mining the potential information association of cross-modal data. Multimodal medical image data analysis aims to improve the expression ability of features by fusing the input data of different modalities according to a specific mechanism to identify the lesion area or stage the lesion accurately. Among them, explaining the principle of multimodal medical image segmentation and building the algorithm through information change is a crucial task of multimodal medical image analysis.
Medical image segmentation is mainly used to extract the contour of interest from medical imaging data. This process performs pixel-wise partitioning by identifying anatomical structures in the image. Multimodal magnetic resonance imaging (MMRI) medical images can help doctors fully grasp the specific characteristics of anatomical information of different brain lesions and improve the segmentation performance. In the segmentation task of abnormal brain lesions, the location distribution, pixel intensity distribution, and lesion shape vary significantly among different samples, which causes problems with the multimodal segmentation task, such as information loss and inaccurate segmentation results. There is redundant information between medical images of different modalities, which will cause the joint learning algorithm to misjudge the importance of different modalities [1], ignore specific modality information and eventually reduce the prediction accuracy and lead to overfitting [2]. Therefore, this paper studies multimodal brain tumor image segmentation methods based on MMRI data. Currently, the standard image segmentation methods based on deep learning, such as the model based on Fully Convolutional Networks for semantic segmentation (FCN), the model based on U-Net, and the model based on Transformer, have gradually become the mainstream methods for multimodal brain tumor image segmentation. The core of these methods is to map the original data into a higher-dimensional feature space through complex, multi-dimensional, nonlinear array operations and to abstract the data into features that are more satisfying for downstream tasks through segmentation network training. The proof of Radmacher’s complexity shows that the quality of feature space directly affects the accuracy and generalization of the model. We expect to construct a feature space as complete as possible through the design of the network structure to maximize the contribution of the feature space to the task. The feature space will provide the information associated with the given task, suppress the information that interferes with the task, and complete a task-oriented data “compression” work to avoid information redundancy and information loss caused by the design of the network structure. However, these segmentation networks cannot accurately classify at the pixel level because the size of the feature map will gradually decrease during the downsampling process, which affects the effect of precise segmentation. For example, the FCN model replaces the convolutional layer with the deconvolution layer to achieve upsampling. However, the deconvolution layer lacks the mechanism of parameter sharing, leading to its failure to capture the global features in the image, thus losing part of the semantic information. The U-Net model captures image features by multiple down-samplings in the encoder, but when the original size of the image is restored by various up-samplings in the decoder, some details and semantic features will be lost, making the segmentation results inaccurate enough. The Transformer model only considers single-scale feature representation, which will cause information loss.
Aiming at the problems of existing medical image segmentation algorithms, this paper proposes a brain tumor MMRI segmentation network based on feature decoupling and information bottleneck theory: FDIBMNet. The main contributions are as follows:
(1) Based on the reversible network, a multi-direction flow and multi-branch encoder with bottom-up and multimodal information interaction is constructed, which allows information to flow across layers among branches, realizes the transfer of low-level detailed information and high-level semantic information between modes, enhances the expression ability of features and the learning ability of the network and helps the network to maintain richer and more comprehensive information and avoid information loss.
(2) To maintain the information balance of multi-directional flow, the feature fusion module is designed, and the interdependence between feature channels is established to realize the adequate flow and fusion of features between different resolutions, maintain their information is not compressed or lost, and form a feature representation without information loss.
(3) Construct a feature space as complete as possible, maximize its contribution and correlation information to the task, and suppress information that has no contribution and correlation to or interferes with the task. Firstly, based on the topological properties of medical images (pixel connectivity and adjacency), a feature decoupling module is designed to decouple the relationship between pixels and geometric properties of medical image features from the feature space and to model and enhance the topological representation between interested pixels. Secondly, based on the information bottleneck theory, loss constraints were applied to multimodal features to ensure a strong correlation between features and tasks and remove redundancy, and third, based on the mutual information constraint between input and output, decoupling the sharing and uniqueness of multimodal features.
This paper is organized as follows: Section 2 introduces related work. Section 3 introduces the network structure of the algorithm constructed in this paper in detail. Section 4 conducts detailed experiments and evaluates the experimental results. Section 5 summarizes the findings, contributions, and limitations of this paper.
The medical image segmentation method based on deep learning, combined with the basic principles of computer vision, can realize the automatic recognition and segmentation of the foreground area of the lesion area and the background area, such as organs and tissues, through the fine pixel classification of the medical image, without the need for manual feature design and extraction. Through convolution and pooling operations, finer high-level semantic features are gradually extracted, making the segmentation results closer to the actual situation. Deep learning technology improves the accuracy and segmentation efficiency of medical image analysis and supports the early detection, diagnosis, and treatment of diseases. Standard supervised learning segmentation algorithms include medical image segmentation algorithms based on the FCN framework, medical image segmentation algorithms based on the U-Net framework, and medical image segmentation algorithms based on Transformer framework.
FCN is a classical image segmentation method. By converting the Convolutional Neural Network (CNN) structure into a fully convolutional structure, the network can output a segmentation result the same size as the input image. FCN utilizes convolutional and deconvolution operations to learn the category label of each pixel in the image to achieve pixel-level segmentation. Sun et al. [3] proposed multi-channel FCN for liver tumor segmentation from CT images. Each stage of contrast-enhanced CT images provides different information about pathological features, so a network is trained for each stage of contrast-enhanced CT images, and their high-level features are fused to achieve automatic liver tumor segmentation of CT images. Ben-Cohen et al. [4] proposed an algorithm for segmenting liver cancer metastases on CT, combining FCN with super-pixel sparse classification to achieve accurate segmentation of small lesions and reduce the false positive rate. Feng et al. [5] proposed a multi-stage FCN architecture for 2D MRI segmentation of the prostate. The algorithm can capture more accurate spatial information and prostate boundaries through different sequences of MRI. Compared with CNN-based image segmentation methods, FCN can accept input images of any size and become more efficient without size transformation. However, FCN does not fully consider the relationship between pixels, and the pixel-level segmentation results still cannot meet the requirements of accurate segmentation. The medical image segmentation method based on the U-Net framework can deal with the above problems of FCN well.
Unlike the segmentation task of natural scene images, medical images usually contain artifacts, and the boundary of the region of interest is blurred and unclear. U-Net can effectively combine low-resolution features with high-resolution features through jump connections, and it has become a benchmark framework for most medical image segmentation tasks. Chen et al. [6] proposed a long-range sensing model for the segmentation of fuzzy boundaries of medical images, which has long-range solid sensing ability and can effectively perceive the semantic context information of the entire image. Experimental results show that the proposed algorithm is more effective in improving the segmentation accuracy of fuzzy boundary regions than other long-distance sensing methods such as Transformer. Yu et al. [7] proposed a differential evolution algorithm based on U-Net for medical image segmentation. The algorithm relies on the expertise of differential evolution to search neural networks automatically. The variable length encoding strategy is used to optimize the neural architecture, effectively improving brain tumors’ segmentation effect. Zhang et al. [8] proposed an algorithm integrating densely connected convolutional modules into the U-Net architecture. By replacing the standard convolutional layer with dense connections, the width of the network is increased, and the features are extracted without increasing the parameters to make the network deeper. In the research of multimodal fusion data analysis algorithms for brain tumors, a downsampling block is used to reduce the size of the feature map to accelerate learning, and an upsampling block is used to adjust the size of the feature map to achieve accurate reconstruction of segmented images. The algorithm achieves precise segmentation of brain tumors on MRI.
U-Net algorithm can obtain high-precision segmentation results in image segmentation tasks of various modalities and diseases, especially when dealing with small targets or images with complex details. U-Net network architecture is relatively simple, easy to implement, and understand. However, the encoder part of U-Net mainly focuses on extracting local information, and it is challenging to integrate global information, which may lead to poor performance in processing long-distance dependency and context information, especially in the task of medical image segmentation. At the same time, during the coding process of U-Net, the down-sampling operation will lose part of the spatial information, which is a challenge for tasks that require accurate spatial localization. The decoder part of U-Net makes it difficult to effectively recover global information during upsampling, which may lead to decreased accuracy of segmentation results. Compared with U-Net, the medical image segmentation algorithm based on the Transformer framework can deal with global information well.
The medical image segmentation algorithm based on the Transformer framework captures the long-distance dependence through the self-attention mechanism and effectively processes the global information in the image. Jiang et al. [9] proposed a label decoupling network with a space-channel graph convolution and a dual attention enhancement mechanism. The algorithm constructs learnable adjacency matrices and uses graph convolution to efficiently capture global long-range information on spatial locations and topological dependencies between different channels in an image. The dual attention enhancement mechanism is constructed, and the edge attention mechanism module is designed in the edge branch to promote the learning ability of spatial region and boundary features. The algorithm can retain the spatial location information in the medical image and improve the accuracy of medical image segmentation. Zhang et al. [10] used spatial pyramid pooling instead of pooling layers as encoders and integrated attention mechanisms to capture complex cross-dimensional interaction information. Extensive experimental results on brain tumor segmentation datasets show that the proposed algorithm performs excellently in medical image segmentation. Li et al. [11] proposed a mutually reinforcing multi-view information model for lung tumor segmentation. The model uses the attention mechanism to enhance node attributes, designs the gated convolution strategy to integrate the enhanced attributes and original features, constructs the learning context of the multi-channel CT, and realizes cross-channel information fusion. Multi-view mutual information is fused through an interactive attention mechanism. Finally, the node embedding, channel context embedding, and original features are adaptively integrated, and the final output is obtained through the segmentation decoder.
Medical image segmentation algorithms based on the Transformer framework can better model the global context to understand the information in complex medical images. The framework is easy to extend and can further improve the effect of medical image segmentation by increasing the number of layers, input images of different sizes, and adjusting the model size. However, Transformer can only consider global information and has a weak ability to extract cross-domain information.
In conclusion, the medical image segmentation method based on the deep learning framework is superior to the traditional medical image segmentation method, but there are still the following problems:
(1) The direction of segmentation will be limited when the algorithm based on the deep learning framework is used for medical image segmentation. For MRI image segmentation with complex boundaries, such as brain tumors, we can modify the direction of segmentation by adjusting the form of the loss function further to improve the accuracy of brain tumor image segmentation.
(2) The core of the neural network is to map the input data into the feature space through complex, multi-dimensional, and nonlinear array operations and perform feature transformation and nonlinear transformation layer by layer. The original data is mapped into a higher dimensional feature space as the hierarchy is gradually deepened. In this feature space, the data is abstracted into semantic features that are more satisfying for the downstream task. The construction of feature space is closely related to the accuracy and generalization ability of the model. This study expects to map data from the original data space to the “representation space” through network design. Through the training of neural networks, it is expected to maximize the information that contributes to the task and is related to the task and suppress the information that does not contribute to the task, is not associated with the task, or has interference with the task. Perform a kind of data “compression” for downstream tasks.
(3) During the construction of neural networks, avoiding the loss of information caused by the design of network structure is necessary. For example, in the shared segmentation network downsampling process, the size of the feature map will gradually decrease, which cannot represent the specific contour of the object and cannot classify the organization of each pixel, so the goal of accurate segmentation cannot be achieved. However, downsampling can reduce the computational complexity of deep networks and is widely used in network structure design. Therefore, it is necessary to balance feature extraction and computational complexity.
Clinically common MRIs usually contain a variety of different sequences, and different sequences are used to examine different anatomical structures, Common sequences include T1-Weighted Imaging (T1WI), T1-weighted Gadolinium enhanced Imaging (T1Gd), T2-Weighted Imaging (T2WI), and Fluid Attenuated Inversion Recovery (FLAIR). The T1WI sequence showed clear anatomical structure and bleeding, and brighter adipose tissue, with relatively few artifacts, but the lesion was not clearly displayed. The contrast between blood vessels and brain tissue is more obvious in T1Gd sequence, so the blood vessels and lesions of the brain can be more clearly displayed. T2WI sequence is contrary to T1WI sequence, which is clearer for lesions and edema, especially for the diagnosis of brain tumors. The FLAIR sequence can be used to determine the edema area around the tumor. Although MRI takes a long time to image, it is widely used for accurate screening of diseases in clinic because it has no radiation and can observe different anatomical structures and tissue information through different sequences.
Fig. 1 shows the design of the feature decoupling and information bottleneck theory image segmentation network (FDIBMNet) framework. The model consists of an encoder, a feature decoupler, and a decoder. In the encoder part,
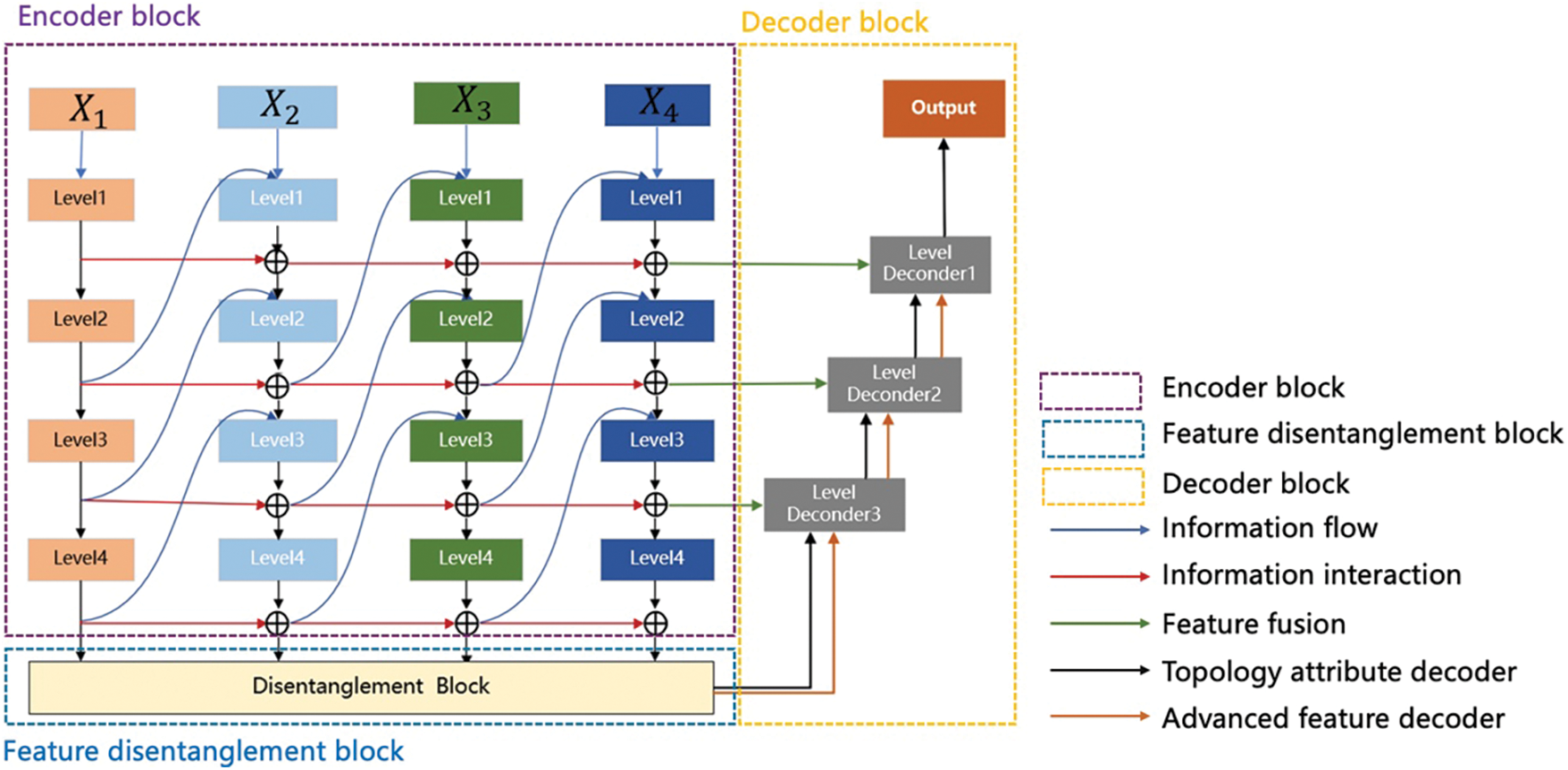
Figure 1: FDIBMNet’s framework
The information between different modalities may have shared or unique information related to the target task, and the information between modalities may have similarities or differences. Only by fully characterizing the information of the modalities can the results of the target task be maximized. Through the interaction of information, the encoder fully represents the modal information and ensures a strong correlation between the information in the feature and the target task. The basic design principle of the feature decoupler comes from the region-growing algorithm. Firstly, the pixel value of the center of the lesion will be given, and the relationship between the surrounding pixels and the pixels of the lesion will further expand the segmented area. When the surrounding pixels are unrelated to the pixels of the lesion area, the boundary delineation of the lesion will be completed. The feature decoupler is set up based on this idea. Through feature coupling and combined with the decoupling directional features, the feature decoupling device ensures the consistency of the anatomical position in the image and completes the extraction of semantic information related to the lesion boundary. In the decoder part, high-level features, topological attributes, and information fusion are used to decode the extracted semantic information and topological characteristics in parallel, which ensures that the structure shape, pixel brightness, texture features, and the relationship between the representation pixel and the surrounding neighborhood are preserved at the same time in the decoding process, and the lesion segmentation is effectively realized.
The encoder comprises four parallel branch encoders containing different levels of feature extraction blocks, and the transmission of lossless information is maintained by a multi-level reversible connection between two adjacent columns. In forward propagation, this architecture scheme can ensure that the information in the model flows in two ways: one is the “top-down” information transmission to achieve single-mode feature extraction, and one is “bottom-up” information transmission to achieve cross-modal and cross-level information interaction. Specifically, multimodal image data
where,
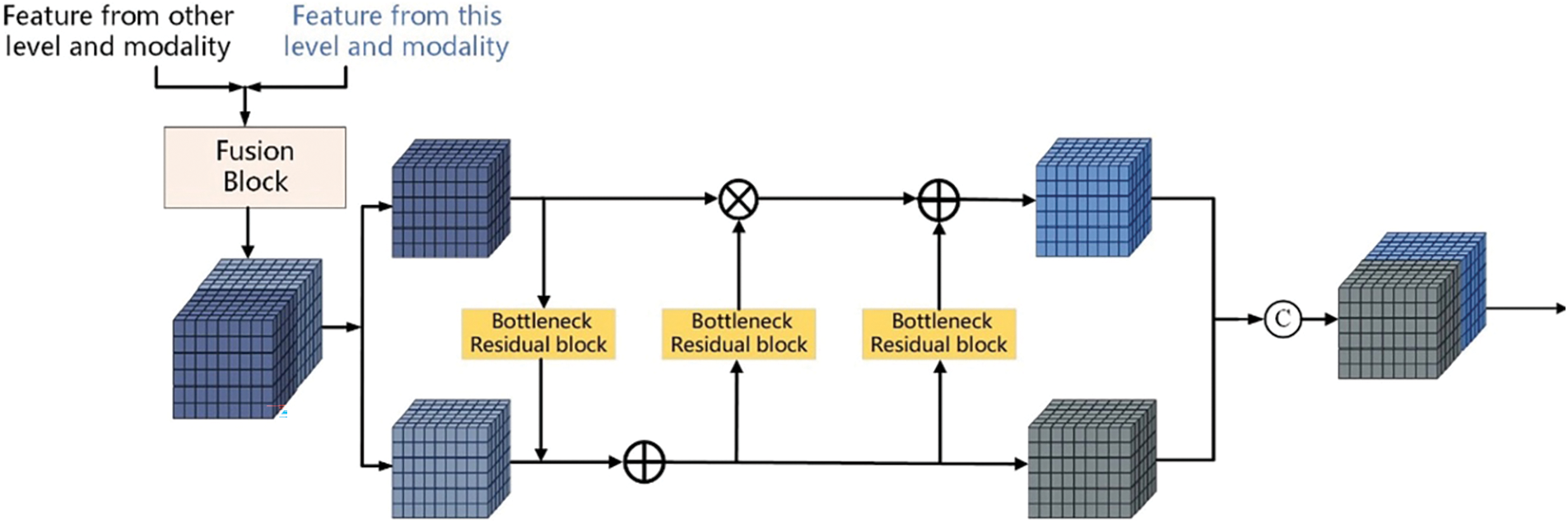
Figure 2: Encoder structure: Level encoder block
This parallel branch structure gradually decouples features during the forward propagation process to maintain the integrity of information and avoid information compression or loss. The bidirectional flow of information between multiple branch networks is allowed so that in the process of forward propagation, each branch structure can receive the information of other branch structures and pass the processed results to the next layer of other branches while retaining all the information in the transmission process.
In forward information propagation, a CNN-based structure is constructed to realize the mapping of input data to features. Ideally, if the extracted features can be “reconstructed” from the input data through some backward process, the information is not lost and is effective in the forward process. Combined with the background of multimodal medical image segmentation in this chapter, it is hoped to design a network structure to ensure the effective extraction and lossless transmission of information such as anatomical characteristics, structure shape, pixel brightness, texture features, and the relationship with surrounding tissues of abnormal brain lesions during the information transformation of input medical images. This is important for later brain lesion tissue segmentation tasks.
Invertible Neural Networks (INNs) is a particular type of neural network whose main feature is that the mapping from input to output is bidirectional and reversible. This property makes the backpropagation and gradient calculation of the network more efficient and reliable and can better achieve data reconstruction and reduction and maintain the integrity of information in the task. INNs are a lossless feature extraction architecture, which is very suitable for information retention and network training in multimodal image fusion. Based on INNs, this paper proposes constructing LB with INNs blocks with affine coupling layers to realize feature extraction and lossless transfer, as shown in Fig. 3a,b [12].
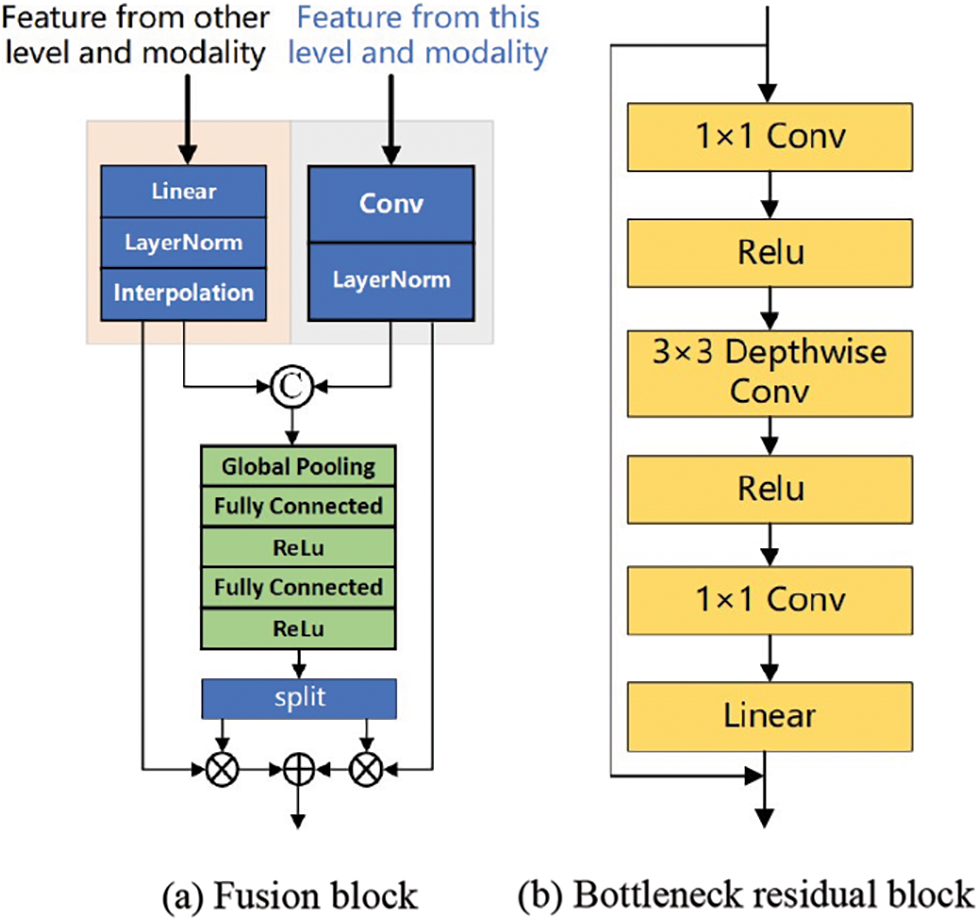
Figure 3: Encoder structure: Fusion block and bottleneck residual block
Firstly, the feature map
where,
The information Bottleneck Residual block (BR) takes the low-dimensional compressed representation as input, first extends it to higher dimensions, and then convolutional layers are used for convolution. Features are subsequently projected back to a low-dimensional representation with linear convolutions. The lossless feature
where,
According to the properties and characteristics of the reversible network, the connection of two side branches can be set to any mapping without affecting the lossless information transmission in this reversible layer. Considering the trade-off between computational consumption and feature extraction power, we adopt BR in MobileNetV2 as the connection of two side branches in the INNs structure. As shown in Fig. 3b, after the features enter the BR, the dimension of the input feature map is increased by 1 × 1 convolution, the number of channels is increased, and the mainstream information is stored in the subspace of the high-dimensional space, which provides the basis for the reversibility of the model. Subsequently, all channels were aggregated by 3 × 3 Depth wise separable convolution (DSC), and the spatial features and channel features were extracted from the information after dimension upgrading. DSC mainly performs group convolution on the feature dimension, first performing an independent depth-by-depth convolution operation on each channel and then performing a 1 × 1 point-by-point convolution operation on all channels. In this process, the number of channels remains unchanged, but the length and width of the feature map become smaller, thereby reducing the parameters required by the convolutional layer and improving the computational efficiency. At the end of BR, the original number of channels is restored by 1 × 1 convolution. This process of first dimension upgrading, DSC convolution processing, and then dimension reduction allows the expressiveness of input and output domains to be decoupled from feature extraction, improves the performance of the model, reduces redundant information, and ensures that the information transmitted through the network can better serve the segmentation task.
Based on INNs, a multi-direction flows multi-branch encoder with bottom-up and inter-modal information interaction is constructed. This design avoids the common information loss problem in traditional deep networks, especially when the network level is profound. Therefore, this design can maintain rich feature representations in deep networks, improving the model’s ability to represent data and learning efficiency. However, the output features of reversible networks mix the useful and useless information of the current task, and it is not easy to achieve a good feature expression ability. Therefore, features need not only to keep the information intact but also to decouple representations.
Feature disentanglement describes separating different features or factors in data representation. In some cases, the input data may contain multiple related but distinct features that may be intricately intertwined in complex ways. The goal of feature decoupling is to decompose these mixed features so that the model can learn the representation of each feature independently to extract more meaningful and practical information. The model can better understand the input data’s intrinsic structure and hidden information with feature decoupling. Feature decoupling usually involves the specific design of the network structure, such as using regularization techniques or adding penalty terms in the loss function to encourage the independence of features. Through feature decoupling, the model is expected to learn more robust and distinguishable feature representations to perform better when facing unknown data. Feature decoupling is a strategy to ensure the quality of features.
Maintaining anatomical consistency in medical image segmentation is essential but extremely challenging, as even minor geometric errors may alter global topological properties and lead to functional errors in downstream clinical decisions. Anatomical consistency in an image can be represented by topological properties, such as pixel connectivity and adjacency [12]. Deep learning-based segmentation methods have made significant progress in capturing inter-pixel dependencies within the latent space of the network using encoder-decoder architectures. Currently, typical segmentation networks model the segmentation problem as a pixel-by-pixel classification task and use segmentation masks as unique labels. However, this pixel-by-pixel modeling scheme is suboptimal because it does not directly exploit inter-pixel relations and geometric properties. Therefore, these models may lead to low spatial coherence in the prediction, that is, inconsistent prediction of neighboring pixels with similar spatial features [13]. When applied to medical data with high noise and artifacts, low spatial consistency may lead to the problem of insufficient extraction of topological attributes. Pixel connectivity has long been used to ensure the fundamental topological duality of separation and connectivity in digital images [14]. For problem modeling, using connectivity masks essentially changes the problem from pixel-by-pixel classification to connectivity prediction, thereby modeling and enhancing the topological representation between pixels of interest. Compared with the segmentation mask, using the connectivity mask as the training label representation provides more information in the following three aspects: first, the connectivity mask stores the classification information between the connections of pixels and has the perception of the relationship between pixels; Second, edge pixels can be represented sparsely [15]; Third, it contains a wealth of directional connection information. Therefore, the latent space constructed by the neural network trained with the connectivity mask has both pixel category features and directional features, and each feature exists in a specific subspace of the latent space. In previous studies [16], these two sets of features are learned simultaneously, maintaining a high coupling in the latent space and introducing redundancy [17]. However, the separation of meaningful subspaces from feature Spaces has been shown to effectively explain the dependence and independence between features. Inspired by feature space decoupling, this study considers decoupling subspaces with different feature meanings from feature space when performing feature extraction and uses the decoupling directional features to enhance the overall data representation for the convenience of subsequent analysis.
The features from the four modalities were first fused. The features
where,
Due to the simultaneous connectivity between category features (whether they belong to a lesion tissue) and orientations in different pixels, it is natural to store orientation information between channels with the deepening of the network. Therefore, each channel of the fused multimodal feature map
As shown in Fig. 4, the weighted excitation
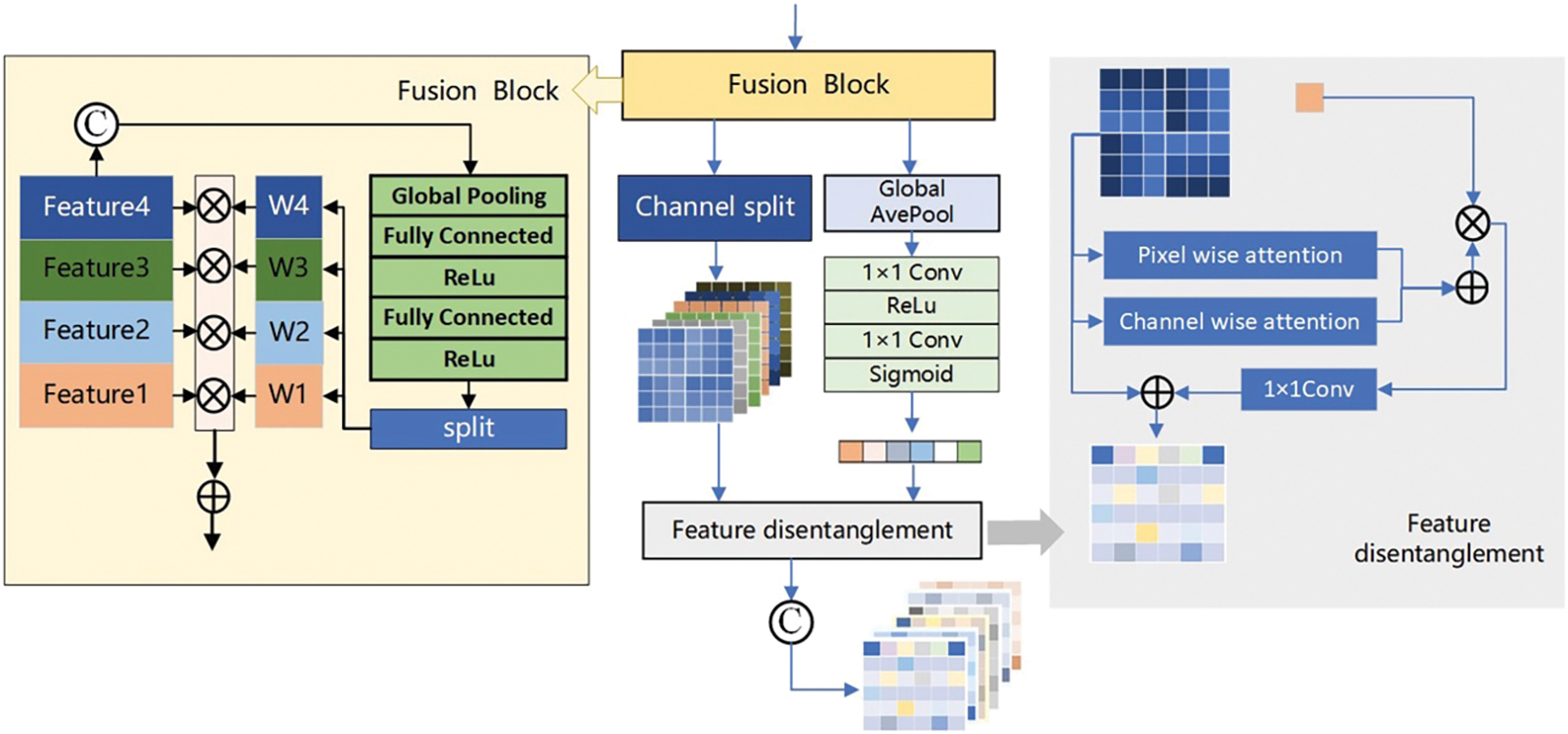
Figure 4: Feature decoupling structure
When medical images pass through each encoder level, the extracted features contain specific topological properties. To ensure that the topological properties of each level are fully utilized, we propose a decoder that can perform feature fusion. As shown in Fig. 5, the input of the decoder is divided into two parts. One part we call high-level features
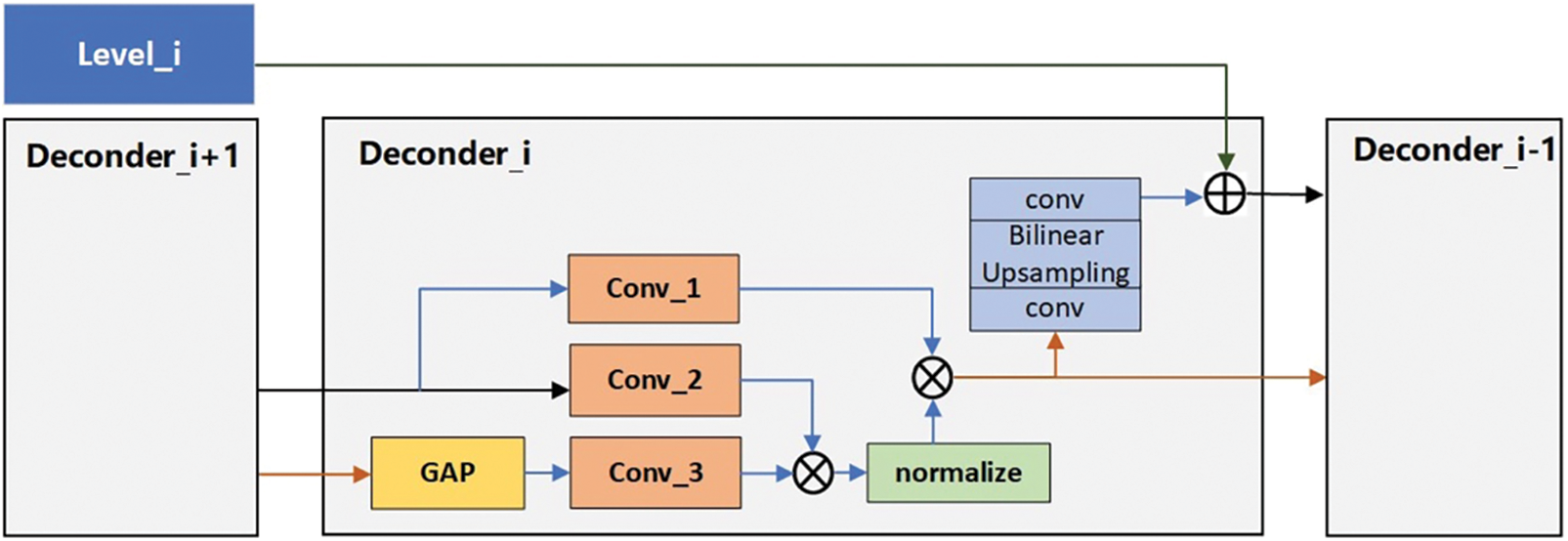
Figure 5: Decoder Structure structure
When
where,
The attention feature map
For medical images, system noise caused by device type and information loss caused by data transmission will cause different degrees of impact on image quality. From the perspective of imaging, although it will not affect human vision in discriminant decision-making, from the perspective of the model, the addition of noise and other disturbances will introduce information that is not related to the decision, affect the data distribution, and then degrade the performance of the model. To avoid the interference of redundant information on the accuracy of the segmentation network, the loss function is constructed based on the information bottleneck theory.
The Information Bottleneck (IB) theory understands the neural network as an encoder and a decoder; the encoder encodes the data into features, and the decoder decodes the features into the output. Its essence maximizes the mutual information between features and production and reduces the information between input and features. To retain the information most relevant to the input and the task. Information bottleneck theory definition:
where,
The experience of Jia et al. [18] shows that using IB as the objective function of Deep Neural Network (DNN) learning will indeed increase the robustness of the model and reduce the sensitivity of feature extraction to input disturbances. However, the calculation of IB is very complex and cumbersome, and researchers use the Hilbert-Schmidt Independence Criterion (HSIC) as a practical computational alternative to IB. HSIC, a statistical dependence measure proposed by Gretton, is the Hilbert-Schmidt norm of the cross-covariance operator between distributions in the reproducing kernel Hilbert space. Like mutual information, HSIC captures the nonlinear dependence between random variables. Loss constraints on multimodal features based on HSIC can ensure a strong correlation between features and tasks and remove redundancy, which is defined as:
where,
where,
Based on the above analysis, we re-examine the HSIC Bottleneck. When extracting cross-modal information, we take it as part of the constraints of multi-modal feature information. Firstly, we ensure that the features extracted from each layer of the network are as highly relevant to the task as possible, and secondly, we ensure that the feature information is unique and shared between modes. Finally, the redundancy of feature information is reduced to ensure that the information contained in the feature map can serve the downstream tasks to the greatest extent, reduce the interference of redundant information, and increase the model’s learning ability and generalization ability. The characteristic loss function based on HSIC Bottleneck is defined as follows:
where, the setting of hyperparameters is empirically selected according to reference, where
Multimodal images are different descriptions of the same target. MRI images with other modalities have different representation advantages for the lesion area, with the uniqueness and sharing of information between modalities. When multimodal data is combined with a specific task, the information contained in the data is first classified: one is the information shared between different modalities related to the task, one is the information that is unique to each modality about the task, and the last is the redundant information that is not associated with the task. Three kinds of information are defined based on mutual information:
(1) All the mutual information between all modal data and tasks.
where,
where, denote all mutual information between joint random variables
(2) Shared and unique information related to modality and task.
where,
where,
In view of this, the multi-modal feature loss function based on mutual information is defined as follows:
Loss constraints on multimodal features decouple the task-related unique information and shared information. The effective representation of multimodal feature information is enhanced by maximizing task-related unique information and shared information and minimizing task-irrelevant redundant information. Starting from this idea, we establish the loss function based on HSIC and mutual information as follows:
where,
This paper was carried out in a hardware environment with an Intel®Core™i9-10900X CPU and an NVIDIA Geforce GTX Titan A100 GPU. The network model is based on the PyTorch framework, Torch version 1.10.2, Cuda version 11.3, and Python version 3.8.10. The model was trained using the Adam optimizer after 300 iterations. The initial learning rate was set to 0.0001, the weight decay to 0.0005, the batch size to 16, and the patch size to
To quantitatively evaluate the proposed method, the Dice coefficient, and HD95 were selected as the evaluation indexes of all segmentation algorithms. Dice is a set similarity measure function commonly used to calculate the similarity between labels and segmentation results. A more considerable Dice value indicates better segmentation. HD95 determines the Hausdorff distance between labels and segmentation results. The smaller the HD95 value, the better the segmentation effect.
The training and testing experiments are carried out on the BraTs2021 dataset to verify the algorithm’s effectiveness proposed in this paper. The effectiveness of the proposed algorithm is further verified on the ANNLIB dataset. Further, to confirm the generalization performance of the proposed algorithm, the trained model is directly used in the test experiment of BraTs2023-MET. Among them, the Tumor segmentation labels of the BraTs2021 dataset included background, Edema (ED), Necrosis and Non-Enhancing Tumor (NCR/NET), and Enhancing Tumor (ET). The purpose of the segmentation task in this chapter is to segment the Whole Tumor (WT), Enhancing Tumor (ET), and Tumor Core (TC) regions.
Ablation studies were performed on the BraTS2021 dataset to evaluate the contribution of the main modules in the methodology. The baseline method is set as follows: the feature decoupling module is replaced by a simple feature stacking method for feature fusion, the decoder uses a simple convolution combination method for feature decoding, and the loss function uses the most straightforward Dice loss and cross-entropy loss weighting. The ablation experiments of four modules (loss function
It can be found from Table 1 that the segmentation results of FDIBMNet reached the optimal in terms of Dice and HD95 in WT, TC, and ET. The effect of the decoder on the whole model was investigated. The ablation experiment showed that the HD95 of WT reached 62.628, the Dice of TC reached 0.829, the Dice of ET was 0.787, and the HD95 was 44.420, all of which achieved suboptimal results. However, although the tumor segmentation effect was practical, it was still not as good as the proposed FDIBMNet model. Considering the impact of
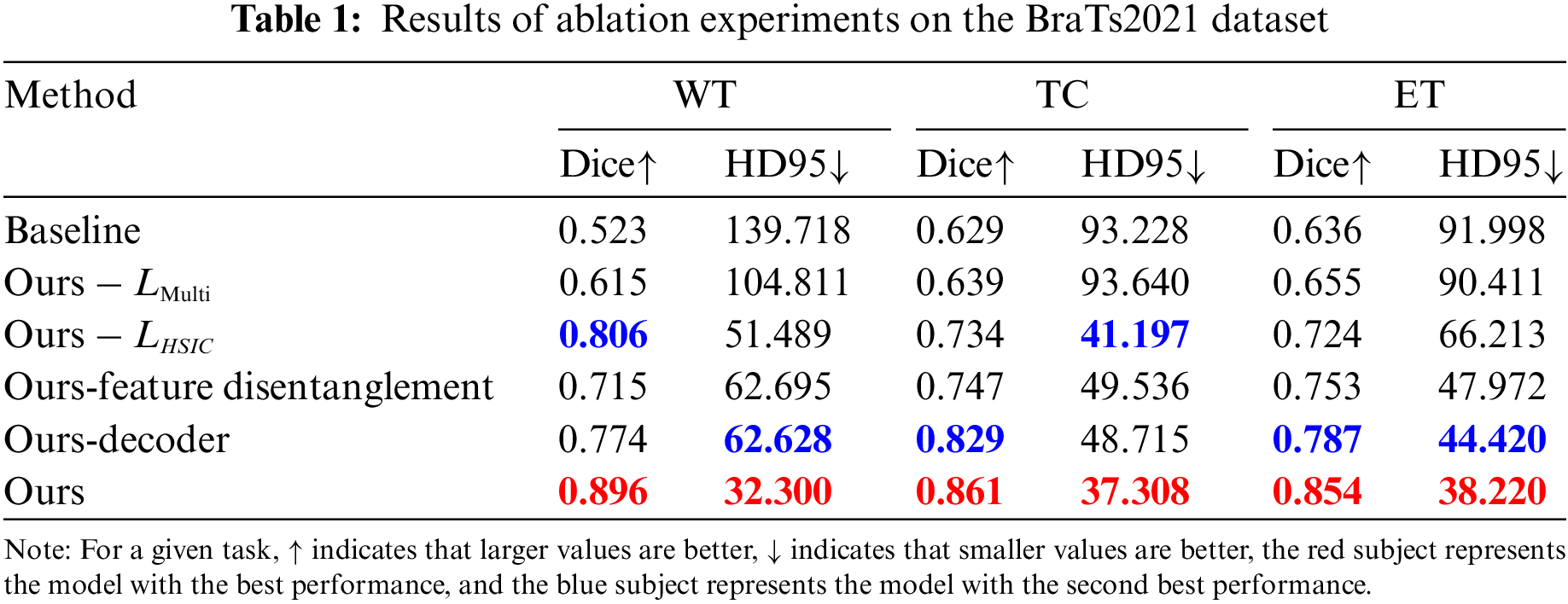
In the feature disentanglement module, the information stacked matrix after the fusion of four modes is input. Through the combination of global pooling, full connection and activation function, the excitation weights of multiple channels in the full mode are obtained, and then the initial dimension reduction work is completed through channel selection and global average pooling. The results were input into the Pixel wise attention and Channel Wise attention of the feature decoupling module to complete the feature decoupling. The results of ablation experiments combined with the network framework to remove the feature decoupler module showed that the Dice of the model decreased by 0.181, 0.082, and 0.101, and the HD95 increased by 30.395, 12.228, and 9.752, respectively. This result indicates that the feature decoupling module can effectively influence the results of the network on the segmentation task. At the same time, to further understand the basic situation of the algorithm, we calculated the number of parameters and floating-point numbers of different structures, as shown in Table 2.

To better prove the convergence of the algorithm proposed in this paper, the epoch of BraTs2021 data set is plotted with the change of loss. Based on Fig. 6, it can be found that after removing the Dice module, the loss function gradually converges when the epoch = 10. After removing the Cross-entropy module, the loss function converges gradually after epoch = 15, and after removing the multimodal fusion module, the loss function converges gradually after epoch = 8. After removing the HSIC module, the loss function gradually converges after the epoch = 9. The convergence of the algorithm is proved.
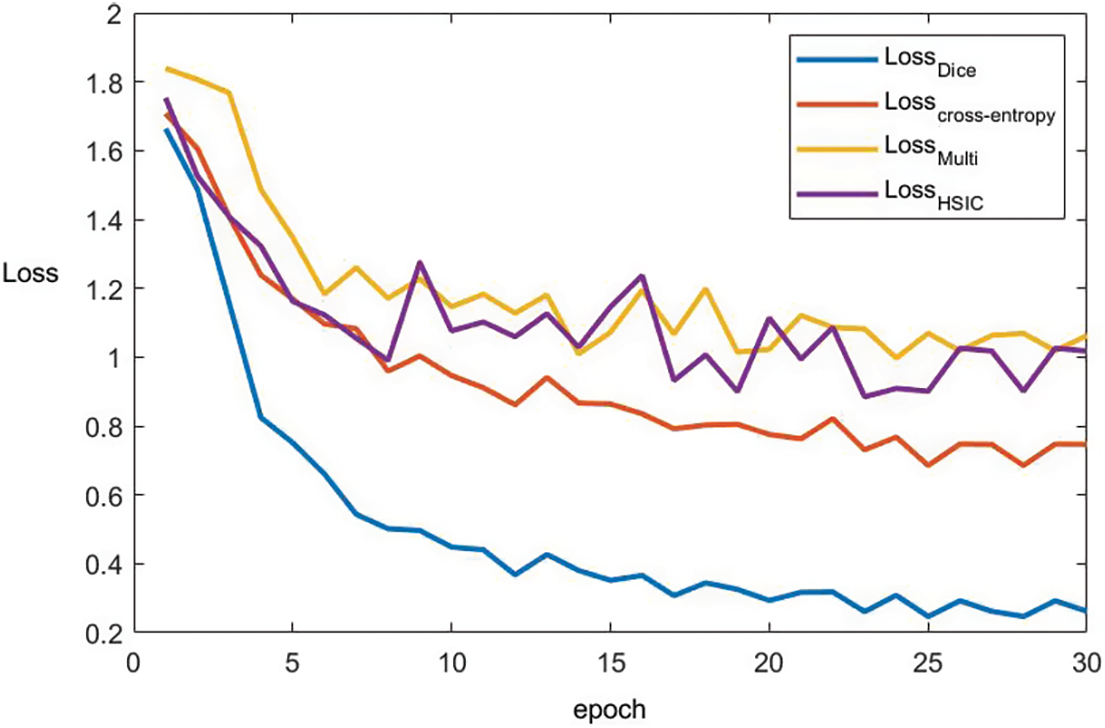
Figure 6: The convergence curve of each loss term
To verify the effectiveness of the proposed method more comprehensively, this paper selects seven segmentation methods that have performed well in the field of multimodal medical image segmentation in recent years for comparative experiments. It includes multimodal medical image segmentation methods based on the CNN framework, multimodal medical image segmentation methods based on the GAN framework, and multimodal medical image segmentation methods based on the Transformer framework Table 3, and the experimental results are presented and analyzed from the perspective of quantitative experiment and qualitative experiment.
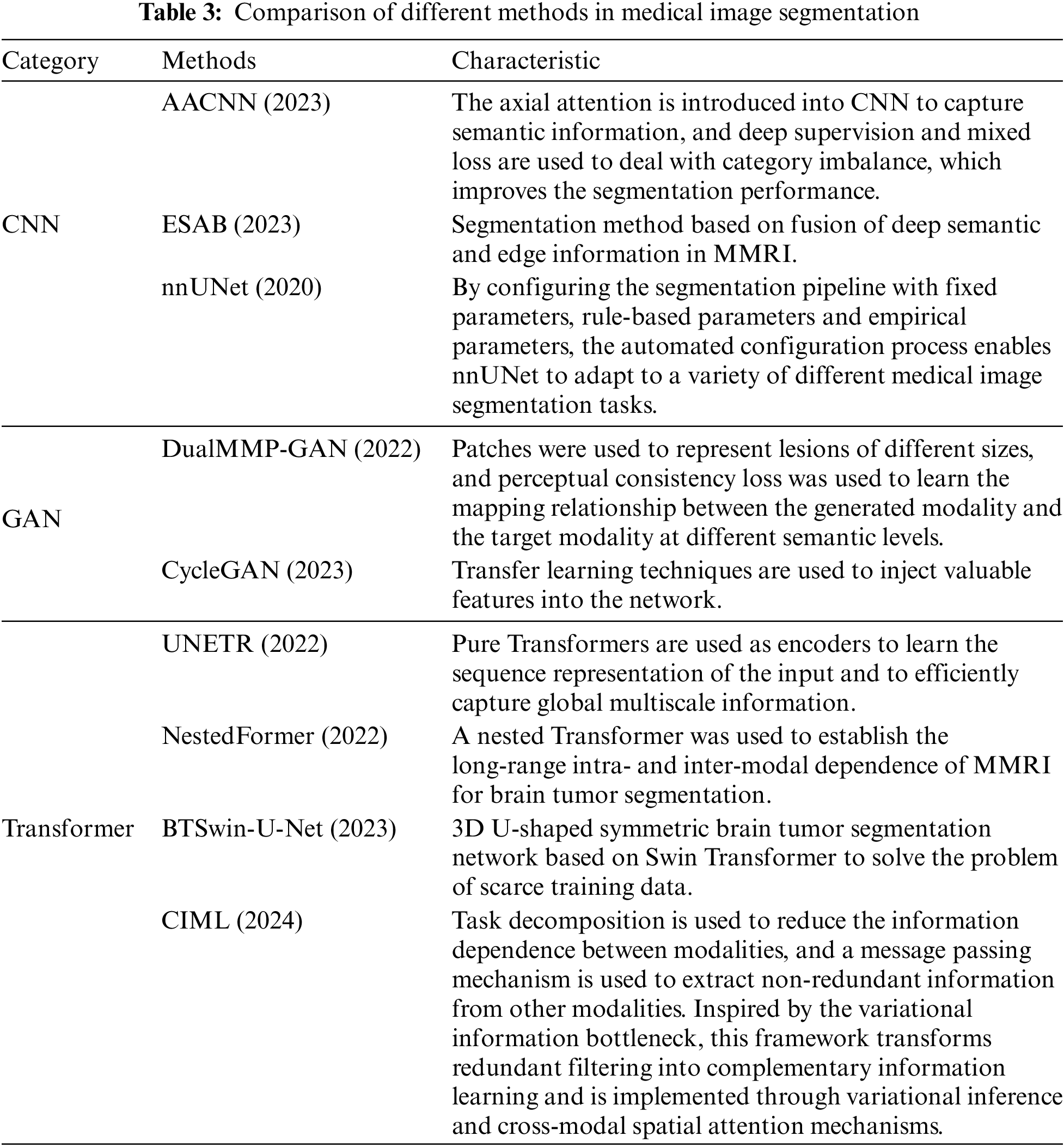
Table 4 reports the experimental results of the seven methods and the method proposed in this chapter on the BraTS2021 dataset under the same experimental environment. FDIBMNet was optimal in Dice and HD95 of WT, TC, and ET. Combining the results of Table 4 and Fig. 7 can be found, compared with the method based on the CNN framework, the Dice of FDIBMNet in WT, TC, and ET were increased by 0.130, 0.074, and 0.083 on average, and the HD95 were decreased by 27.980, 20.323, and 31.252 on average. Compared with the CNN framework, the information flow and feature fusion of FDIBMNet ensure the richness of information in the downsampling step, which can improve the accuracy of image segmentation and reduce the distance between the target region and the segmentation region. Compared with the method based on the GAN framework, the Dice of FDIBMNet in WT, TC, and ET were increased by 0.115, 0.108, and 0.105 on average, and the HD95 decreased by 17.451, 31.486, and 33.933 on average. Compared with the GAN framework, the feature decoupling and topological attribute features of FDIBMNet ensure the quality of the information in the upsampling step, which is conducive to mapping the segmentation results back to the source image. Compared with the method based on the Transformer framework, the Dice of FDIBMNet in WT, TC, and ET were increased by 0.113, 0.080, and 0.071 on average, and the HD95 were decreased by 23.123, 18.557, and 23.065 on average. Like the Transformer framework, FDIBMNet captures multi-scale features and solves the feature relationships within and between modes through information flow. The difference is that FDIBMNet adopts a parallel structure in the down-sampling step, which retains topological attributes and high-level features. This is conducive to improving the effect of medical image segmentation. According to the results of the quantitative analysis, FDIBMNet is significantly better than CNN, GAN, and Transformer in terms of the segmentation effect of WT, TC, and ET of brain tumors.
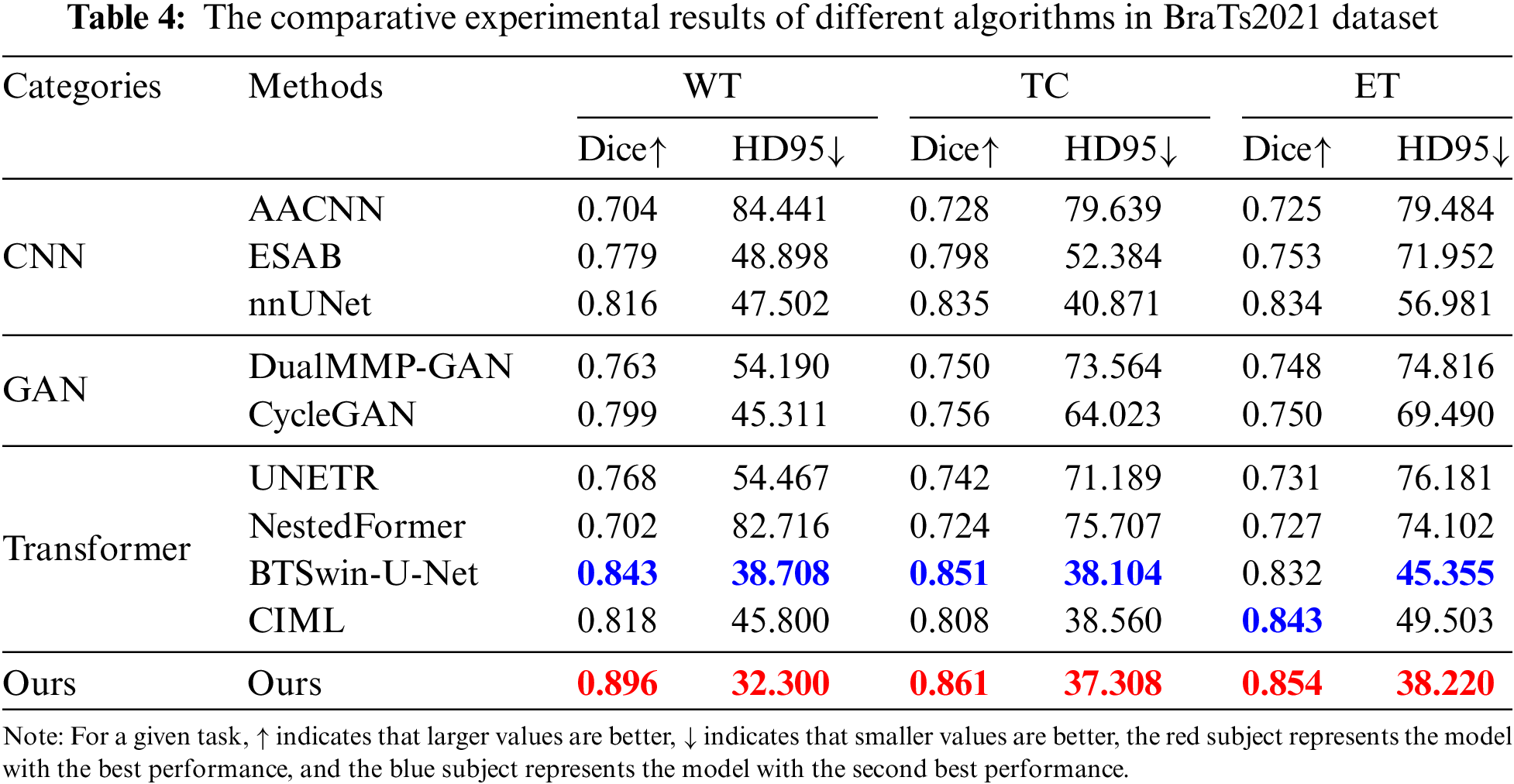
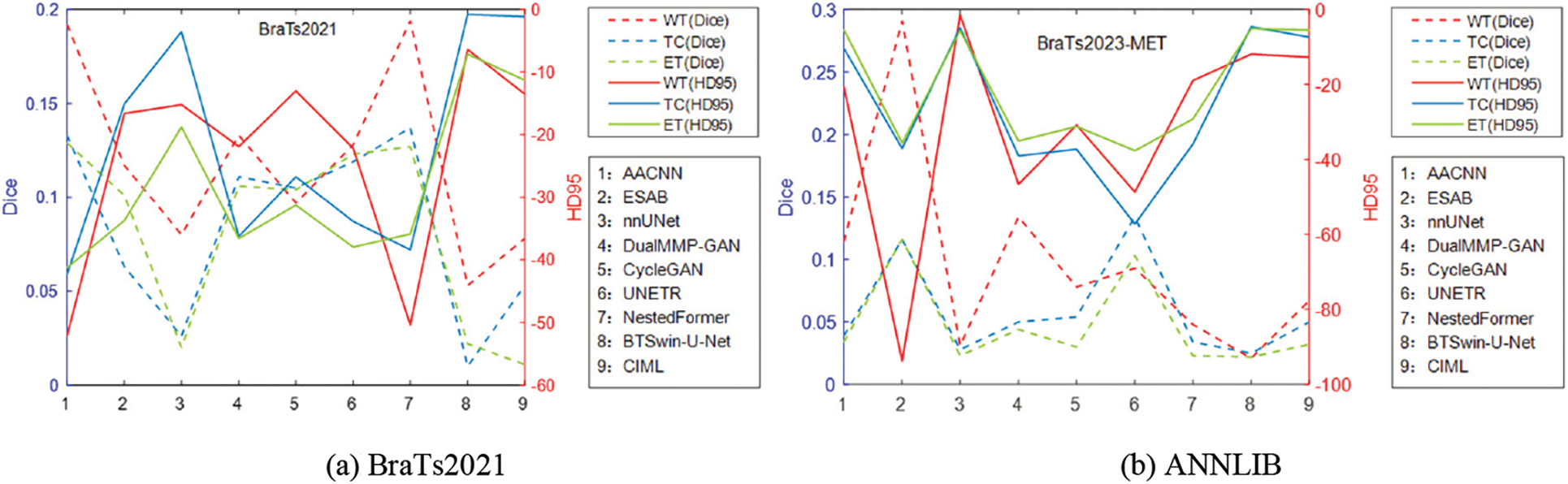
Figure 7: Line plot of the error of BraTs2021 and ANNLIB results
Qualitative analysis was performed using visualization to judge FDIBMNet’s effect further. In this chapter, two sample images are selected to visually compare the impact of different methods on the segmentation task. Fig. 8 shows the segmentation results of brain tumor slices, labels, and techniques. In the source image, the brain tumor was not significantly imaged in the T1WI mode, and there was no noticeable pixel difference from the surrounding tissue. In the FLAIR mode, the edema area was very significant and had a clear boundary with the surrounding tissue, but the tumor core and the enhanced tumor pixel area were concealed, which was challenging to distinguish significantly. In T1Gd mode, only the tumor core and enhanced tumor showed pixel intensity that differed from other tissues, and there was a clear boundary between them. The appearance of the whole tumor was clear on T2WI.
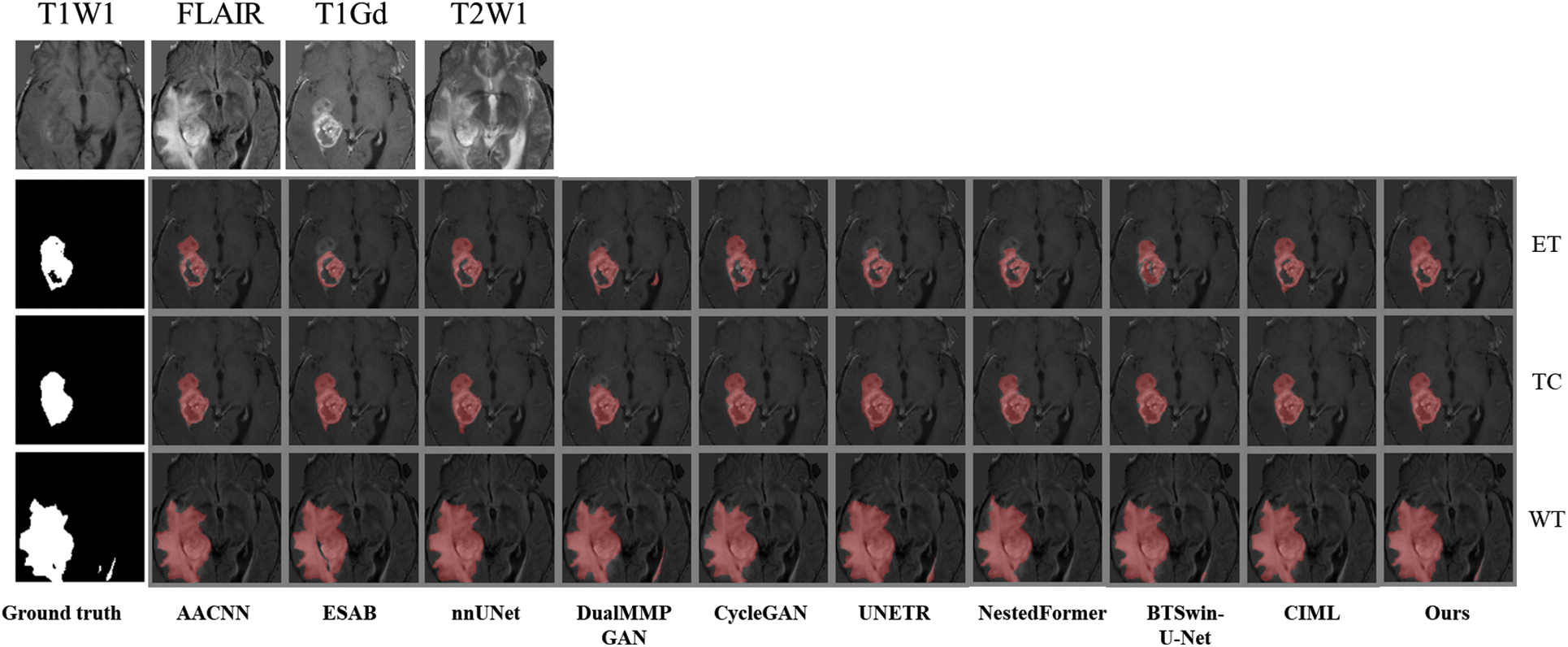
Figure 8: The first example picture contrast method visualizes the results
Combined with Fig. 9, it is found that the medical image segmentation method based on the CNN framework has a poor processing effect on the incoherent lesion area. For example, there are non-tumor areas inside the enhanced tumor area of the first example, and the segmentation effect of the CNN framework on this part is poor, with the phenomenon of under segmentation. At the same time, the boundary segmentation of the tumor core and all tumors was not precise. From the perspective of the network framework, the two methods based on CNN did not enhance the boundary information, resulting in unclear boundaries. The medical image segmentation method based on the GAN framework has a better processing effect on incoherent regions than CNN. The main reason for this phenomenon is that the GAN-based method can extract semantic information at different scales, ensuring the multi-scale richness of information, but a certain degree of boundary still needs to be added. Medical image segmentation methods based on the Transformer framework have poor processing effects on incoherent regions and boundaries. For example, the NestedFormer and BTSvin-U-Net methods in the second example think that there are non-tumor regions in the center of the tumor. The UNETR method showed poor segmentation of the left lower area of all tumors. Although the medical image segmentation method based on the Transformer framework considers the multi-scale information within modalities and the spatial information between modalities, its loss function does not constrain the information. It cannot achieve the effect of accurate segmentation. The FDIBMNet method proposed in this chapter can accurately segment the non-tumor regions inside the tumor and segment the boundaries of the cancer with a good segmentation effect.
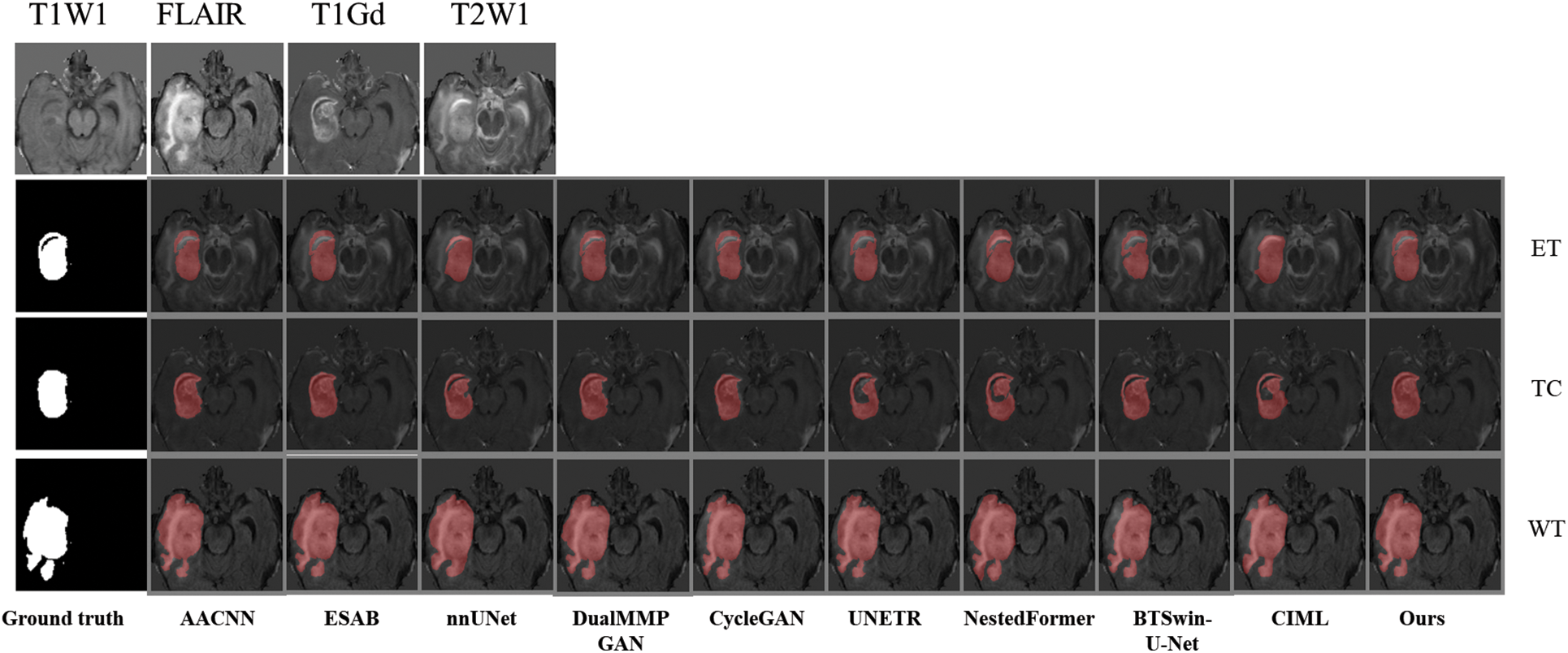
Figure 9: The second example picture contrast method visualizes the results
To better verify the effectiveness of the algorithm, ANNLIB dataset was used to further verify the effectiveness of the algorithm. According to the results in Table 5, it can be found that our proposed method achieves the optimum in both Dice and HD95 indices, in which nnUNet achieves the second-best. Combined with the line chart results, the Dice index was increased by 0.098 on average, and the HD95 index was decreased by 37.936 on average, which further verified the effectiveness of the FDIBMNet method.
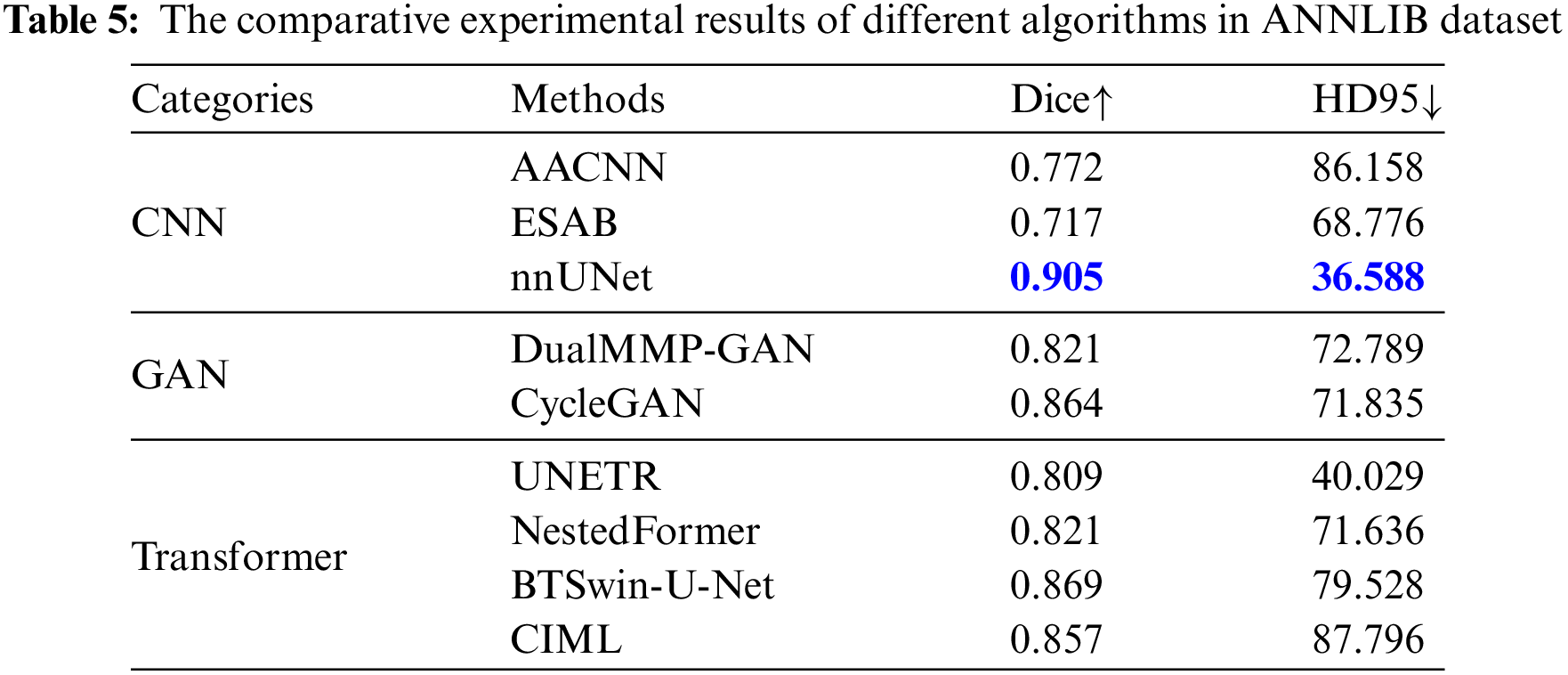
The experimental results of qualitative analysis and quantitative analysis show that the CNN-based image segmentation method could be more friendly to boundary segmentation. The image segmentation method based on the Transformer will segment the tumor area into non-tumor areas prone to undersegmentation. The image segmentation method based on GAN has a better effect than CNN and Transformer, but there is still a certain degree of clarity. Due to the feature decoupling module, the FDIBMNet method proposed in this paper extracts the topological attribute features, ensuring incoherent lesion regions’ segmentation effect. In addition, information flow and fusion combined with mutual information loss function safeguard the richness of information and the extraction of crucial details, thereby improving the performance of the overall segmentation model.
4.4 Generalization Experiments
The trained model based on the BraTS2021 glioma dataset was directly used in the brain metastases segmentation test task of the BraTS2023-MET dataset to verify the generalization of the method proposed in this chapter. The experimental results of Dice and HD95 for all network methods are listed in Table 6 and Fig. 10.
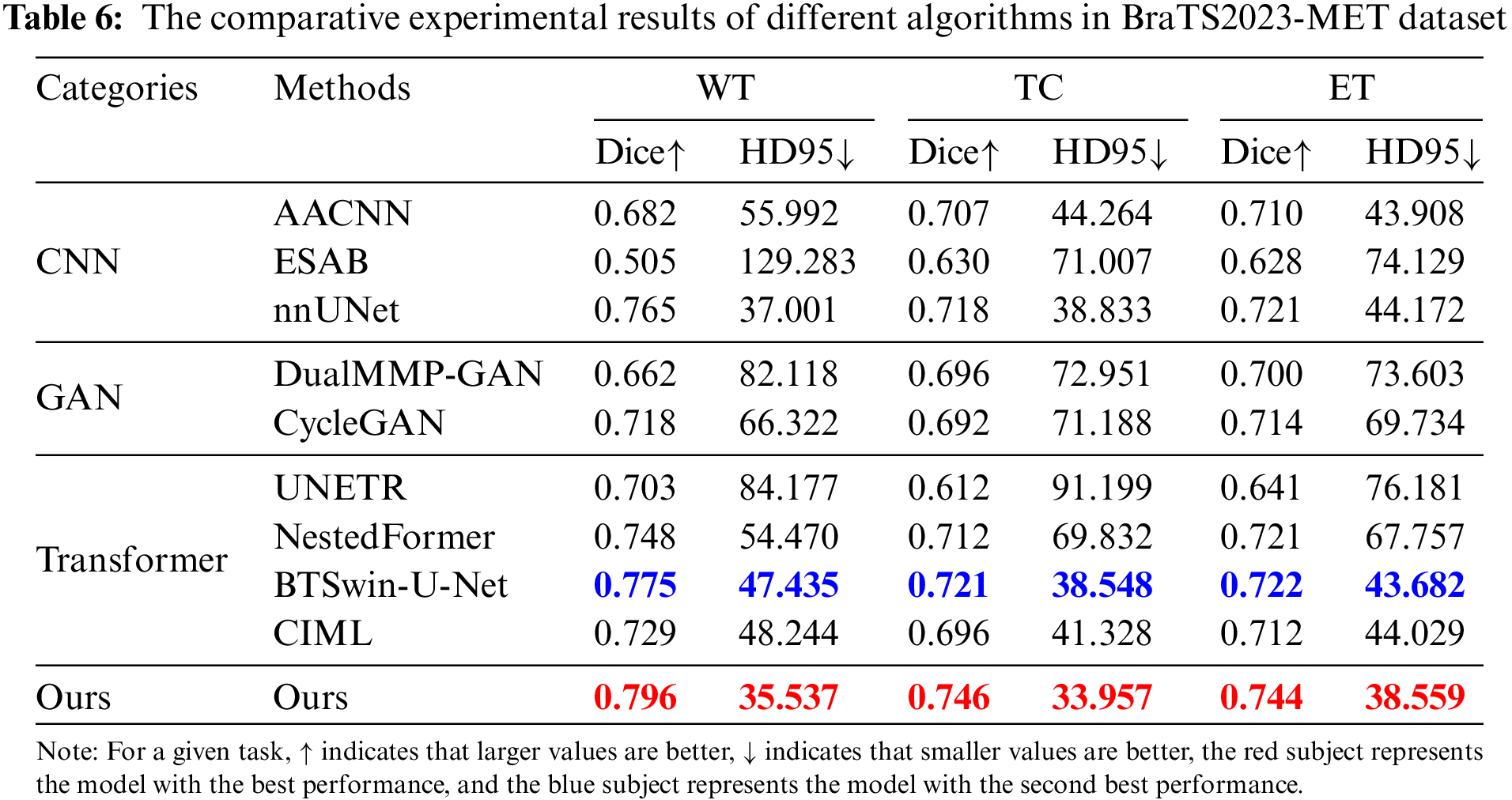
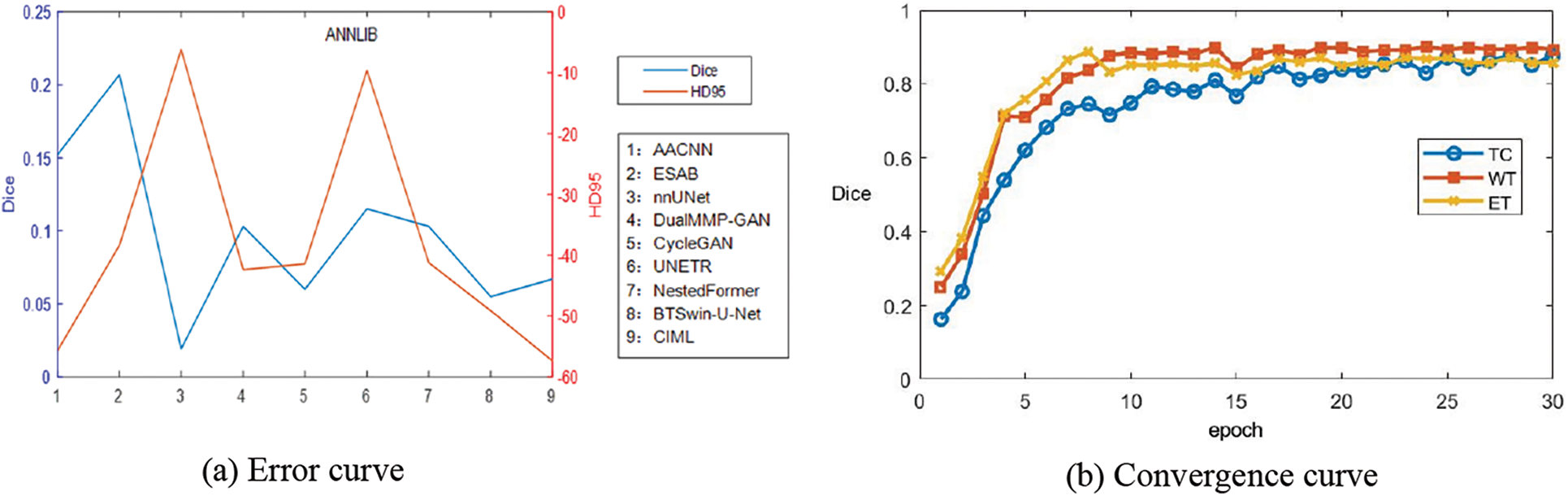
Figure 10: Line plot of the error of BraTS2023-MET results and the convergence curve of Dice of WT, TC, WT
Table 6 and Fig. 10 report the experimental results obtained by the abovementioned methods on the BraTS2023-MET dataset under the same experimental environment. The Dice index of FDIBMNet in the three WT regions, TC regions, and ET regions was 0.796, 0.746, and 0.744, respectively, and the HD95 index was 35.537, 33.957, and 38.559, respectively, which were all optimal values. Compared with the method based on the CNN framework, the Dice scores of FDIBMNet in WT, TC, and ET were increased by 0.145, 0.061, and 0.058 on average, and the Dice scores of HD95 were decreased by 38.555, 17.411, and 15.511 on average, with the most significant performance improvement. Compared with the method based on the GAN framework, the Dice of FDIBMNet in WT, TC, and ET were increased by 0.106, 0.052, and 0.037 on average, and the HD95 were decreased by 38.683, 38.113, and 33.110 on average. Compared with the method under the Transformer framework, the Dice of FDIBMNet in WT, TC, and ET were increased by 0.057, 0.061, and 0.045 on average, and the HD95 were decreased by 23.045, 26.270, and 19.353 on average. On the BraTs2021 dataset, the GAN-based image segmentation method is significantly better than the CNN-based image segmentation method, and the opposite is observed on the BraTS2023-MET dataset. The reason is that the T1Gd modality is missing in the generalization dataset, so the information on the lesion is relatively less, which is not enough to describe the disease state. On the BraTs2021 dataset and BraTS2023-MET dataset, the FDIBMNet method showed the best performance, and this result further verified the segmentation performance of the FDIBMNet method.
To better prove the generalization of the algorithm proposed in this paper, a graph of the change of epoch with Dice of the BraTS2023-MET dataset is drawn as Fig. 10b. Based on the graph, it can be found that when the TC, WT, and ET regions are divided, the Dice gradually converges when the epoch = 15. The generalization of the algorithm is proved.
Qualitative analysis was performed using visualization to judge the generalization effect of FDIBMNet further. In this section, an example image is selected to visually compare the impact of different methods on the segmentation task. According to the visualization results Fig. 11, ESAB and UNETR judged the brighter pixel area in the imaging as the lesion area. Methods BTSLOU-U-NET classifies all tumors with dark pixels as usual, and most of the pixels with strong pixels as tumors, which will lead to misdiagnosis or missed diagnosis. However, the WT region, TC region, and ET region can be segmented relatively entirely by the method proposed in this paper, and the normal tissue is rarely misclassified as the lesion area, reducing misdiagnosis. In particular, the segmentation effect is suitable for regions with blurred edges and strong pixels, showing good anti-interference ability. This is because FDIBMNet emphasizes the decoupling of multimodal information, directional information, and category information so that the extracted information can be directed to the lesion area with direction guidance and reduce the interference of pixel light and dark. In addition to inaccurate brain tumor detection, the ET and TC regions should be included in the WT region, but the BTWCN-U-NET method shows that the WT region does not contain the TC region. The visualization experimental results of the method proposed in this chapter show that FDIBMNet can effectively capture robust correlation information with the help of the
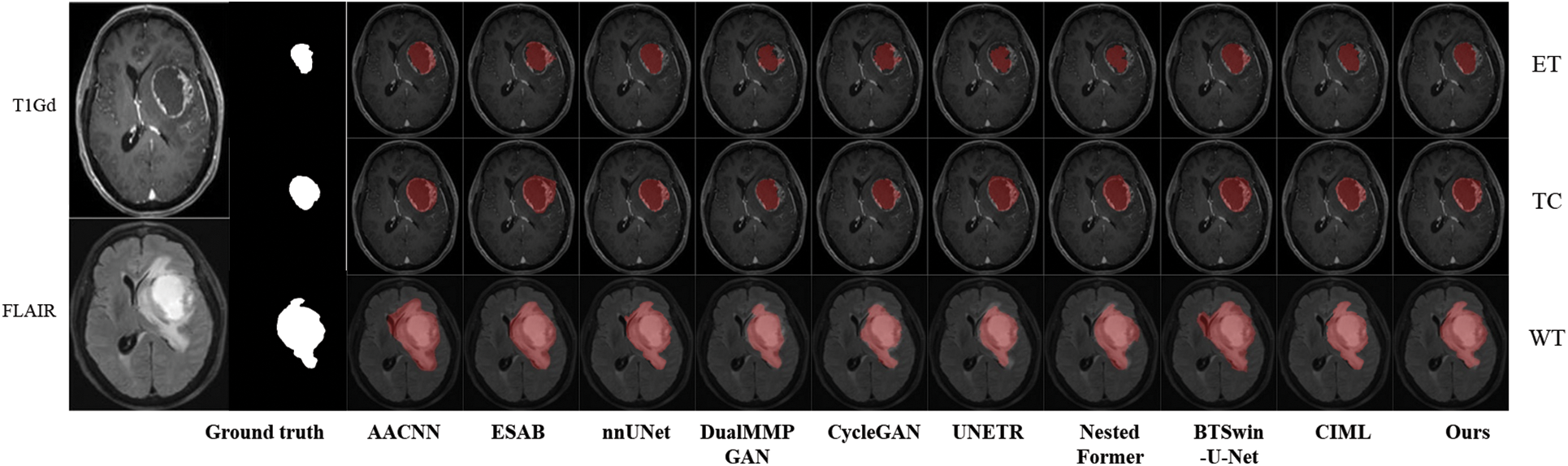
Figure 11: Generalization of experimental results
By combining the quantitative and qualitative analysis of the generalization experiment, FDIBMNet flows the information between different modes through the bottom-up flow, combines the mutual information and the information bottleneck loss function, and ensures the adequate flow of information and improves the quality of feature extraction. The feature decoupling method can effectively obtain high-level features and topological attributes of image segmentation, and with the parallel upsampling structure, the accurate segmentation of incoherent lesion regions and lesion boundaries can be ensured.
We drew the pre-experiment loss function curve to determine the number of training rounds and found that FDIBMNet began to decline smoothly when Epoch was 50 and became stable around Epoch 100. Therefore, Epoch 100 was selected for formal experiments in this paper (Fig. 12).
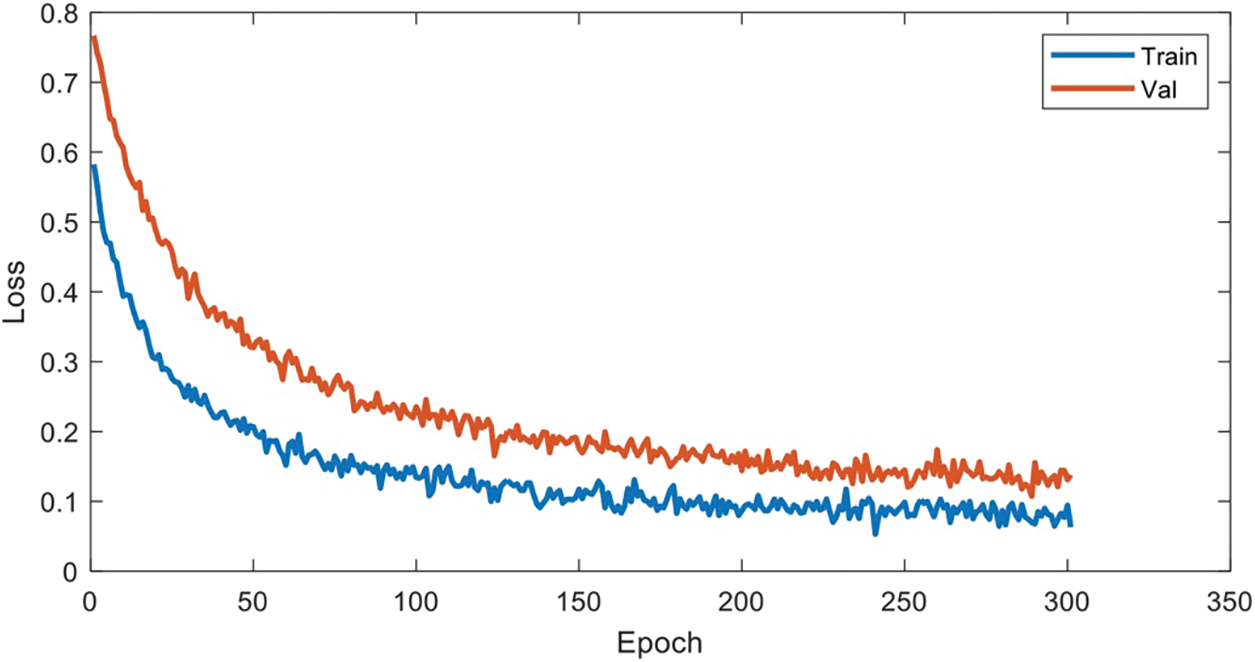
Figure 12: Hyperparametric analysis
This paper proposes a novel multimodal segmentation framework for brain tumors based on feature decoupling and information bottleneck theory (FDIBMNet). The proposed framework contains a multi-branch encoding structure, a feature decoupler, and a two-branch decoding structure. Firstly, based on INNs, a multi-direction flow and multi-branch encoder with bottom-up and inter-modal information interaction was constructed, and a “complete” “representation space” was built to maximize the information that contributed and correlated to the task and suppress the information that did not contribute and correlated to the task or interfered with the task. Secondly, based on the topological properties of medical images (pixel connectivity and adjacency), a feature decoupling module was designed to decouple the relationship between pixels and geometric attributes of medical image features from the feature space and to model and enhance the topological attribute representation between interested pixels. From the information bottleneck, loss constraints are applied to multimodal features to ensure a strong correlation between features and tasks and remove redundancy. Finally, based on the mutual information constraint between the input and output, the sharing and uniqueness of multimodal features are decoupled, and the effectiveness and generalization of the proposed segmentation method are verified on BraTS2020 and BraTS2023-MET datasets. As a visualization task of segmentation results, this paper can combine image fusion and image classification tasks to form a multimodal and multi-task medical image auxiliary diagnosis system for brain tumors. Accurate brain tumor segmentation can help doctors better understand the location, size and morphology of tumors, to develop more precise treatment plans and reduce the damage to healthy brain tissue. In addition, multimodal methods can integrate information from different imaging modalities, such as structural information, functional information and metabolic information, to provide a more comprehensive description of tumor characteristics, which is of great significance for personalized medicine and precision medicine. Therefore, the research in this paper not only promotes the application of machine learning technology in the field of medical image processing, but also provides technical support for improving the efficacy and safety of brain tumor treatment. But there are still some shortcomings in this paper. With the continuous development of multimodal fusion technology, the multi-modal image fusion segmentation technology can be studied from the perspective of diffusion model. It is also possible to consider extending to large model directions.
Acknowledgement: We thank the National Natural Science Foundation of China for supporting this project. We would like to thank Li Yang for the guidance of this article. Thanks to Shanghai Zhangjiang Institute of Mathematics for the support of computing power.
Funding Statement: The manuscript was supported by the following grants: Beijing Natural Science Foundation (No. Z210003); National Natural Science Foundation of China (NSFC12026607); National Natural Science Foundation of China (NSFC12031016); Key R&D Program of the Scientific Research Department (2020YFA0712203); Key R&D Program of the Scientific Research Department (2020YFA0712201).
Author Contributions: Xuemei Yang: Data curation, Methodology, Software, Validation, Writing—original draft and review. Yuting Zhou: Conceptualization, Supervision, Writing—review and editing. Shiqi Liu: Data curation, Investigation, Visualization. Junping Yin: Conceptualization, Funding acquisition, Methodology, Resources, Supervision, Writing—review & editing. All authors reviewed the results and approved the final version of the manuscript.
Availability of Data and Materials: This article uses the public data sets are as follows: BraTS2021 (https://www.synapse.org/Synapse:syn25829067/wiki/610863) (accessed on 05 December 2024), BraTs2023-MET (https://www.synapse.org/Synapse:syn51156910/wiki/622553) (accessed on 05 December 2024), ANNLIB (https://www.med.harvard.edu/AANLIB/home.html) (accessed on 05 December 2024).
Ethics Approval: The relevant data ethics work in this paper has been approved by the Chinese Academy of Engineering Physics.
Conflicts of Interest: The authors declare no conflicts of interest to report regarding the present study.
References
1. X. Li et al., “Adversarial multimodal representation learning for click-through rate prediction,” in Proc. Web Conf. 2020 (WWW’20), New York, NY, USA, Association for Computing Machinery, 2020, pp. 827–836. [Google Scholar]
2. S. Mai, Y. Zeng, and H. Hu, “Multimodal information bottleneck: Learning minimal sufficient unimodal and multimodal representations,” IEEE Trans. Multimed., vol. 25, pp. 4121–4134, 2023. [Google Scholar]
3. C. Sun et al., “Automatic segmentation of liver tumors from multiphase contrast-enhanced CT images based on FCNs,” Mach. Learn. Graph. Anal. Computat. Biomed., vol. 183, no. 3, pp. 58–66, 2017. doi: 10.1016/j.artmed.2017.03.008. [Google Scholar] [PubMed] [CrossRef]
4. A. Ben-Cohen, E. Klang, A. Kerpel, E. Konen, M. M. Amitai and H. Greenspan, “Fully convolutional network and sparsity-based dictionary learning for liver lesion detection in examinations,” Neurocomputing, vol. 275, no. 9, pp. 1585–1594, 2018. doi: 10.1016/j.neucom.2017.10.001. [Google Scholar] [CrossRef]
5. Y. Feng et al., “Multi-stage fully convolutional network for precise prostate segmentation in ultrasound images,” Biocyber. Biomed. Eng., vol. 43, no. 3, pp. 586–602, 2023. doi: 10.1016/j.bbe.2023.08.002. [Google Scholar] [CrossRef]
6. S. Chen et al., “LD-UNet: A long-distance perceptual model for segmentation of blurred boundaries in medical images,” Comput. Biol. Med., vol. 171, no. 3, pp. 108–120, 2024. doi: 10.1016/j.compbiomed.2024.108120. [Google Scholar] [PubMed] [CrossRef]
7. C. Yu, Y. Wang, C. Tang, W. Feng, and J. Lv, “EU-Net: Automatic U-Net neural architecture search with differential evolutionary algorithm for medical image segmentation,” Comput. Biol. Med., vol. 167, no. 10, 2023, Art. no. 107579. doi: 10.1016/j.compbiomed.2023.107579. [Google Scholar] [PubMed] [CrossRef]
8. Z. Zhang, C. Wu, S. Coleman, and D. Kerr, “Dense-inception U-Net for medical image segmentation,” Comput. Methods Programs Biomed., vol. 192, no. 10, 2020, Art. no. 105395. doi: 10.1016/j.cmpb.2020.105395. [Google Scholar] [PubMed] [CrossRef]
9. Q. Jiang, H. Ye, B. Yang, and F. Cao, “Label-decoupled medical image segmentation with spatial-channel graph convolution and dual attention enhancement,” IEEE J. Biomed. Health Inform., vol. 28, no. 5, pp. 2830–2841, 2024. doi: 10.1109/JBHI.2024.3367756. [Google Scholar] [PubMed] [CrossRef]
10. Z. Zhang, Y. Wen, X. Zhang, and Q. Ma, “CI-UNet: Melding convnext and cross dimensional attention for robust medical image segmentation,” Biomed. Eng. Lett., vol. 14, no. 2, pp. 341–353, 2024. doi: 10.1007/s13534-023-00341-4. [Google Scholar] [PubMed] [CrossRef]
11. Y. Li, Y. Wu, M. Huang, Y. Zhang, and Z. Bai, “Attention-guided multi-scale learning network for automatic prostate and tumor segmentation on MRI,” Comput. Biol. Med., vol. 165, no. 1, 2023, Art. no. 107374. doi: 10.1016/j.compbiomed.2023.107374. [Google Scholar] [PubMed] [CrossRef]
12. H. Liu, M. Shao, Y. Qiao, Y. Wan, and D. Meng, “Unpaired image super-resolution using a lightweight invertible neural network,” Pattern Recognit., vol. 144, no. 2, 2023, Art. no. 109822. doi: 10.1016/j.patcog.2023.109822. [Google Scholar] [CrossRef]
13. L. Zhang, B. Verma, D. Stockwell, and S. Chowdhury, “Density weighted connectivity of grass pixels in image frames for biomass estimation,” Expert. Syst. Appl., vol. 101, no. 7, pp. 213–227, 2018. doi: 10.1016/j.eswa.2018.01.055. [Google Scholar] [CrossRef]
14. X. Wang, H. Liu, J. Zhu, Y. Sheng, and C. Zhang, “Deep multimodal medical image fusion network based on high and low frequency feature decomposition,” J. Graph., vol. 45, no. 1, pp. 65–77, 2024. [Google Scholar]
15. A. Rosenfeld and R. Klette, “Digital geometry,” Inf. Sci., vol. 148, no. 1, pp. 123–127, 2002. doi: 10.1016/S0020-0255(02)00284-0. [Google Scholar] [CrossRef]
16. Z. Yang, S. Soltanian-Zadeh, and S. Farsiu, “BiconNet: An edge-preserved connectivity-based approach for salient object detection,” Pattern Recognit., vol. 121, no. 12, 2022, Art. no. 108231. doi: 10.1016/j.patcog.2021.108231. [Google Scholar] [PubMed] [CrossRef]
17. M. Kampffmeyer, N. Dong, X. Liang, Y. Zhang, and E. P. Xing, “ConnNet: A long-range relation-aware pixel-connectivity network for salient segmentation,” IEEE Trans. Image Process., vol. 28, no. 5, pp. 2518–2529, 2019. doi: 10.1109/TIP.2018.2886997. [Google Scholar] [PubMed] [CrossRef]
18. Y. Jia, M. Salzmann, and T. Darrell, “Factorized latent spaces with structured sparsity,” in Adv. Neur. Inf. Process. Syst. 23: 24th Annu. Conf. Neural Inf. Process. Syst. 2010, Vancouver, BC, Canada, 2010. [Google Scholar]
Cite This Article
 Copyright © 2025 The Author(s). Published by Tech Science Press.
Copyright © 2025 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools