 Open Access
Open Access
REVIEW
Gut microbiome modulation: Ancillary effects of inorganic nanoparticles on gut microflora
1 Department of Biological Sciences, International Islamic University, Islamabad, 44000, Pakistan
2 Department of Nanomedicine, California Innovations Corporation, San Diego, CA, 92037, USA
3 Drug Delivery and Cosmetics Lab (DDCL), GCPS, Faculty of Pharmacy, Gomal University, Dera Ismail Khan, 29050, Pakistan
* Corresponding Authors: BUSHRA UZAIR. Email: ; BARKAT A. KHAN. Email:
BIOCELL 2023, 47(2), 245-260. https://doi.org/10.32604/biocell.2023.025311
Received 05 July 2022; Accepted 28 August 2022; Issue published 18 November 2022
Abstract
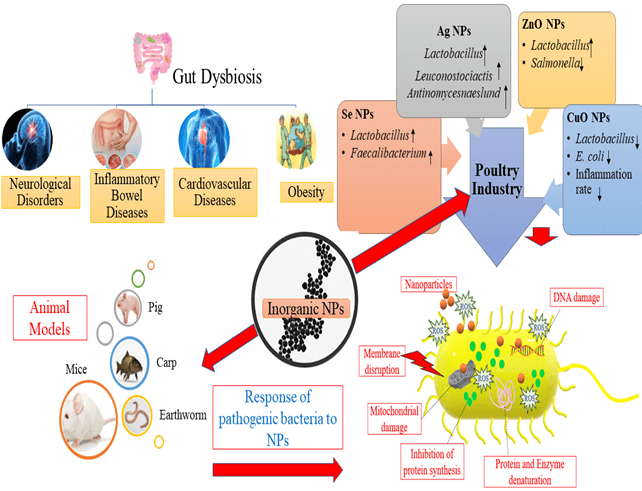
The association of gut microflora and human health is being increasingly recognized, and the impact of gut microflora on the host is well characterized, including the body’s energy metabolism and immune system maintenance. Several human diseases, including metabolic, autoimmune, obesity, hypothyroidism, and intestinal disorders, are closely associated with gut dysbiosis. Inorganic nanoparticles (NPs) are extensively utilized in numerous fields due to their distinctive, attractive physicochemical properties. Estimation of the potential impacts of NPs, with a high number of microorganisms inside the human body (microbiota) and its genomes (microbiome), represents one of the most important aspects of nano-toxicology. This review article aims to provide information on the association of gut microflora alterations to diseases and describe the impacts of various inorganic NPs, including silver, zinc, selenium, titania, silicon, and copper, on gut microflora. Research on the effect of inorganic NPs on gut microflora of animal models and the poultry industry is reviewed. The response of pathogenic Enterobacter species to inorganic NPs has been expounded in detail. This review also highlights the need to focus on the ancillary effects of various inorganic NPs on gut microflora to expedite the suitable advancement of these particles for future use. Finally, the key opportunistic areas for the application of nanotechnology are underlined to manipulate the microbiome of gut dysbiosis, provide an overview, and address potential challenges and our perspective on this evolving field.Graphic Abstract
Keywords
The human microbiome comprises nearly 10–100 trillion symbiotic microorganisms, which comprise the human microbiome, which also includes the genes these microbial cells harbor (Ursell et al., 2012). From the results of current epidemiological, physiological, and quantification studies, complemented by cellular research and animal experiments, it seems that microbial communities may intervene or alter a significant portion of the environmental impacts on human health and the risk of disease (Lynch and Pedersen, 2016). These microbial communities, collectively known as microbiota, comprise a large number of interacting microbes such as fungi, archaea, bacteria, eukaryotic viruses, and bacteriophages concomitantly present on the surface of the human body and in all body cavities (Lynch and Pedersen, 2016). These can be categorized as commensal or mutualistic microbes (Lynch and Pedersen, 2016). Most of the microbes that make up the human microbiota reside in the human intestine and are influenced by the mode of birth, newborn feeding, drug intake, lifestyle, and genetic makeup of the host. The gut microbiota contributes considerably to host immunity training, digestion of food, gut endocrine function regulation, neurological signaling, metabolism of drugs and activity modification, toxins removal, and generating several compounds that impact the host (Fan and Pedersen, 2020; Khan et al., 2021).
Gut microbiota changes are often linked with diseases and are described as gut dysbiosis. Often the compositional change and the associated phenotype may alter a small number of microbes. Gut dysbiosis is linked with numerous intestinal and extra-intestinal diseases comprising infections, asthma, obesity, neurological disorders, colorectal cancer (Schwabe and Jobin, 2013), inflammatory bowel disease (IBD) (Buttó and Haller, 2016; Kamada et al., 2013), allergies and diabetes (Gevers et al., 2014; Hill et al., 2012). Relative microbial abundance variations are caused by dietary changes, infection, immune dysfunction, inflammation, or toxin or antibiotic exposure (Zeng et al., 2017). Various approaches to modify the gut microbiome, including antibiotics, probiotics, prebiotics, symbiotic, post-biotic, dietary changes, and microbiota transplants, lack specificity to achieve targeted changes, so novel approaches are required for gut microbiome intervention aimed at different disease therapies to navigate the complex microecology that exactly intervene with the liable pathways (Song et al., 2019). Nanotechnology can meet the requirement to connect via molecular and macroscopic length scales and is crucial for interaction with insignificant molecule metabolites, microscopic bacteria, and macroscopic structures (Song et al., 2019). Nanoparticles (NPs) can enter the gastrointestinal tract (GIT), making it important to appropriately investigate the potential toxicity risks (Li et al., 2019).
Herein, we review the properties of healthy intestinal microbiota and intestinal dysbiosis leading to other disorders and the role of NPs in solving this problem and also describe the attributes of different metallic NPs that are specifically suited to intestinal microbiome intervention. The toxicity of NPs against gut pathogenic microbes, including Citrobacter rodentium, Salmonella enterica serovar Typhimurium, Shigella flexneri, C. difficile, and Listeria monocytogenes is highlighted by describing their antimicrobial mechanism of action. We highlight critical work in which inorganic NPs have been used to maintain gut homeostasis. “Opportunities to the nascent field of nanotechnology, major challenges to using them as food for gut homeostasis, and some future perceptions to fill the research gaps in this area are elaborated.”
Gut microbiota-homeostasis vs. dysbiosis
A significant assumption for arguing the interruption of the gut microflora is to provide knowledge of the composition and role of the gut microbiome of healthy individuals (Fan and Pedersen, 2020). It is usually subjugated by anaerobic bacteria, which outnumber 100- to 1000-fold aerobic and facultative anaerobic bacteria. Intestinal microbiota as a whole have been reported to comprise 500–1000 species that belong to the few known phyla; among these, two dominating phyla in the gut are Firmicutes and Bacteriodetes, and some other species belong to phyla Proteobacteria, Verrumicrobia, Actinobacteria, Fusobacteria and Cyanobacteria (Huttenhower et al., 2012; Qin et al., 2010). Recently, a large-scale study explored nearly 35000 species of bacteria residing in the human gut (Jandhyala et al., 2015). The relative distribution of these microbial species is distinctive to an organism, partly due to variations in microbial growth rates and structural variants at strain levels within the microbial genes (Huttenhower et al., 2012; Korem et al., 2015; Zeevi et al., 2019) and partly due to the influence of substantial inter-individual variations in environmental exposures and host genetics (Rothschild et al., 2018). Healthy gut microbiota is characterized by constant microbiome functional cores, microbial gene strength, and high taxa variety (Huttenhower et al., 2012). Microbial richness is affected by intestinal transit; thus, only gut bacterial variations and richness are not neutral markers of a healthy microbiome (Falony et al., 2018). Long transit time can contribute to enhanced richness, but not definitely in a healthy intestinal microflora (Fan and Pedersen, 2020). The gut bacteria are important for numerous features of the host biology; as they are involved in the growth and differentiation of colonic epithelium and immune system of host, they are capable of metabolizing the indigestible polysaccharides and production of essential vitamins and provide protection against opportunistic pathogens (Sommer and Bäckhed, 2013). Eventually, for a variety of vital functions, the host relies on its gut microflora, so this gut microflora may contribute to health.
Gut microbiota is fluctuated by the exposure to numerous environmental factors like drugs, toxins, pathogens, psychological and physiological stress, and diet as well (Sommer and Bäckhed, 2013). The composition of the gut microflora is altered by intercurrent infections and treatment by antibiotics (Jandhyala et al., 2015).
People in developed countries exhibit more decline in microbial diversity as compared to those living traditional lifestyles, and have never been exposed to antimicrobial drugs of the modern world (Clemente et al., 2015). The abundance of Bacteroides, Prevotella, Desulfovibrio, Lactobacillus, and Oxalobacter genera in the gut microflora is deteriorating in tandem with urbanization, sewerage, superior housing standards, and better sanitation in general (Clemente et al., 2015; Sommer and Bäckhed, 2013; Tyakht et al., 2013). This decrease in diversity leads to an increase in the incidence of common diseases. Likewise, low levels of microbes are linked with insulin resistance, an increase in adiposity, dyslipidemia, and inflammation in both lean and obese individuals (Le Chatelier et al., 2013). Antibiotics to fight infectious diseases can contribute a key role in the decline of gut microflora; like the use of antibiotics before or during pregnancy and in early childhood can alter the gut microflora makeup in children and infants, ultimately leading to the early onset of obesity (Fujisaka et al., 2016). So abnormal gut microflora is closely associated with neurological disorders, metabolism in obesity, cardiovascular disorders, hypothyroidism, inflammatory bowel diseases, and cancer. Fig. 1 represents the impact of various factors on gut dysbiosis, including diet, drug intake, sedentary lifestyle, pathogen exposure, and stress, demonstrating the disturbance in gut microflora.
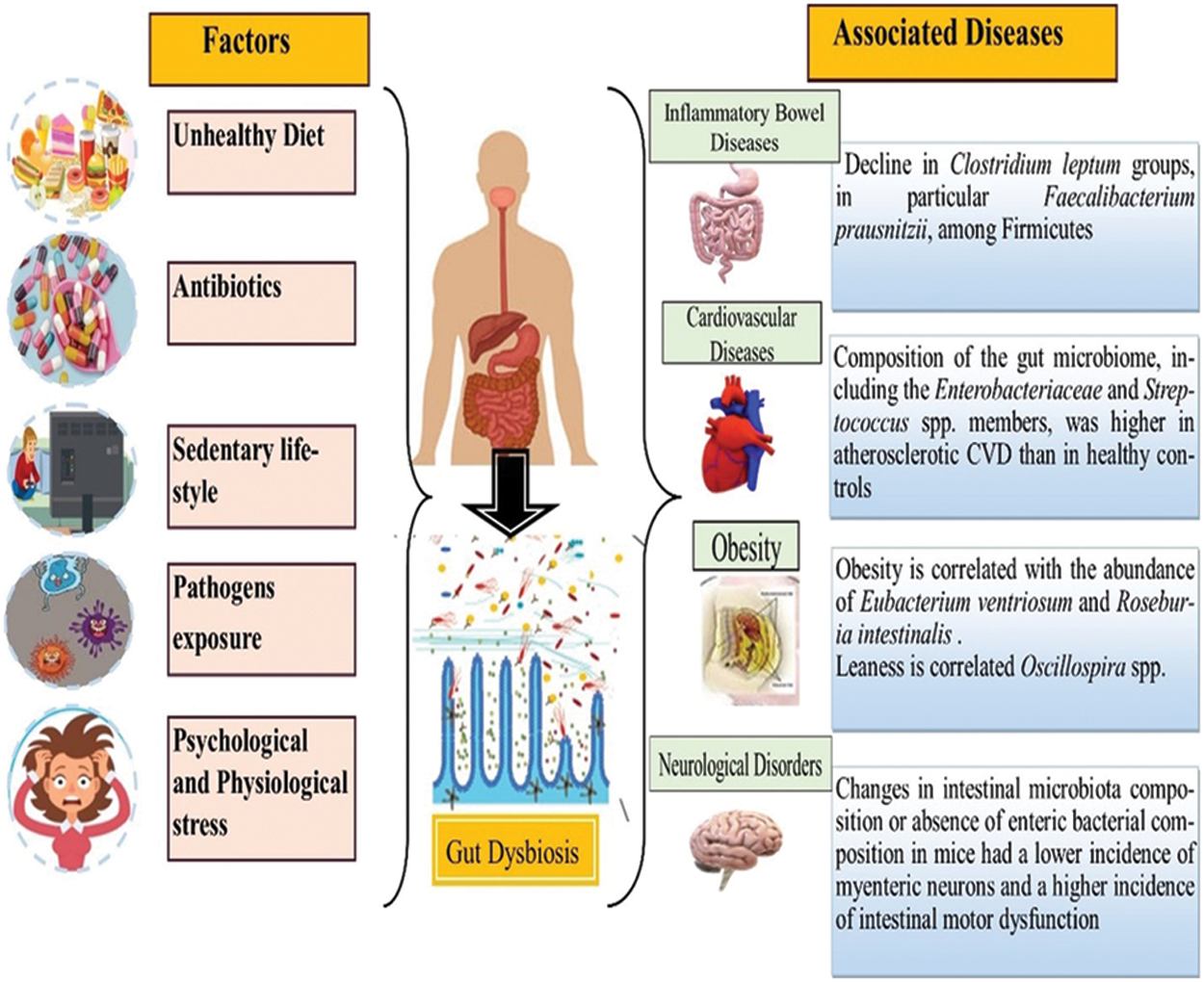
Figure 1: Gut dysbiosis with associated disorders: inflammatory bowel disease, cardiovascular diseases, obesity, and neurological disorders caused by various factors including unhealthy diet, antibiotics, sedentary lifestyle, pathogenic exposure, psychological and physiological stress that disturb the normal gut microflora.
Impact of heavy metal Cadmium (Cd) on gut microbiota
Cd is a non-essential heavy metal designated as a highly toxic pollutant for human health (Kumar and Sharma, 2019). Ingestion of food and drinking water contaminated with Cd increases the metal load in the body. Previous reports have shown the harmful effects of Cd compounds in aquatic and terrestrial wildlife like crabs, fish, zebra, and rodents (Gonzalez et al., 2006; Sun et al., 2018; Wang et al., 2011b). Depending on the concentration and exposure duration of Cd, varied health effects have also been observed in humans. Acute ingestion can have primary effects, such as gastrointestinal (GI) disturbances like diarrhea, vomiting, nausea, and abdominal pain. Chronic exposure can increase the risk of chronic lung, and kidney dysfunction, reproduction impairment, and bone deformation and can also lead to the progression of cancerous cells (Bist and Choudhary, 2022).
Cd has toxic effects on microbial growth by disturbing the protein synthesis and functions of varied enzymatic systems. Gut microbiota is considered the primary contact for Cd and can have extremely deleterious effects on the gut microflora composition. A study reported by Zhai et al. (2017) demonstrated the specific and time-dependent alterations in the gut microflora of mice after exposure to Cd and other toxic metals. Fazeli et al. (2011) reported that Gram-positive bacteria in the gut are more sensitive than Gram-negative bacteria to Cd. Similarly, in the gut, Bifidobacteria were more sensitive to Cd toxicity than Lactobacillus because Lactobacilli were more abundant, with a greater tolerance to Cd than Bifidobacteria. Lactobacillus also has massive Cd binding and removal capabilities (Duan et al., 2020).
Liu et al. (2020) compared the effects of Cd exposure on normal and antibiotic-treated mice that showed that Cd-induced gut dysbiosis could increase intestinal permeability. Kim et al. (2015) reported a weaker proinflammatory response to Cd metal exposure in germ-free mice than that in control mice, indicating that the Cd-induced proinflammatory effect was partially dependent on the gut microflora. Transplantation of gut microbiota from Cd-treated mice into Cd unexposed mice prompted inflammatory and allergic responses in the Cd unexposed mice (Kim et al., 2017). Ba et al. (2017) reported Cd-induced changes in young male mice with an enhanced ratio of Bacteroidetes/Firmicutes and body fat. Transplantation of fecal microbiota from Cd-exposed mice to control mice exhibited increased accumulation of fat; treatment of these mice with oral antibiotics significantly prevented this tendency, indicating the important role of gut microflora in this process (Duan et al., 2020). These findings will help to determine how the gut microflora exerts its impact on the health of the host. Hence, gut microflora is critical for determining the toxicity of environmental contaminants, specifically trace heavy metals like Cd.
Influence of Gut Microflora on Health and Diseases
Since the mid-twentieth century, the prevalence of obesity and its metabolic co-morbidities have increased significantly in developed countries (World Health Organization) (World Health Organization, 2000). According to the World Health Organization, more than 1900 million people above 18 years of age had a body mass index (BMI) of more than 25 kg/m2, and 600 million people were categorized as obese with a BMI of more than 30 kg/m2 in 2016 (World Health Organization) (World Health Organization, 2017). Enhanced consumption of food and a sedentary lifestyle with a widespread phylogenetic susceptibility are the main reasons for the obesity epidemic (McAllister et al., 2009), which is further compounded by the extensive use of antibiotics (Cox and Blaser, 2015).
Obesity may promote the development of the metabolic syndrome and type 2 diabetes with other related comorbidities. In the last 10 years, these metabolic changes have been found to be closely related to gut dysbiosis. Alterations in the gut microflora are associated with the obesity and its related disorders (Angelakis et al., 2012). Moreover, evidence for the function of gut microflora in arbitrating some of the environmental consequences of obesity pathogenesis is accumulating. Successive epidemiological findings have indicated variations in the gut microflora of organisms with obesity and lean organisms. In 2006, researchers found that a versatile obesity-associated microflora would trigger weight increase in lean mice (Turnbaugh et al., 2006). Twin studies at the species level discovered that obesity is correlated with the large number of Eubacterium ventriosum and Roseburia intestinalis, which are short chain fatty acids (SCFA) producers (Tims et al., 2013), while leanness is correlated with butyrate producers such as Oscillospira spp (Gophna et al., 2017) and the methanogenic archaeon Methanobrevibacter smithii (Miller et al., 1982). In another metagenome-wide association investigation, the abundance of Bacteroides thetaiotaomicron, a glutamate-fermenting commensal, was distinctly reduced in people with obesity and was contrarywise associated with serum glutamate intensity (Liu et al., 2017). The bacterial component of gut microflora is less diverse in obese patients as compared to the eutrophic subjects. In a research study, obese mice were observed with increased Firmicutes and decreased Bacteroidetes in feces irrespective of the diet intake (Le Chatelier et al., 2013; Ley et al., 2006). An increase in the Firmicutes/Bacteroidetes ratio was also observed in the fecal microflora of obese humans. However, a recent meta-analysis did not confirm the lower ratio of Bacteroidetes in obese patients and thus suggests that the alterations in gut microflora by weight or BMI may not be universally true (Angelakis et al., 2012). Investigations of intestinal microbial paths and gene families indicate that obesity is related to a reduced unidirectional conjugation ability that transmits genetic information between bacteria and a decline in superoxide reductase, possibly contributing to intestinal oxidative stress (Thingholm et al., 2019).
People with obesity possess higher levels of SCFA and decreased residual food calories in feces as compared to lean individuals. In spite of all accumulated evidences, there still exist knowledge gaps on the role of gut microflora in the development of obesity and how its management may help to control the syndrome.
Gut microflora in inflammatory bowel disease (IBD)
IBD is a disorder described as persistent and deteriorating inflammation of the intestine, and two clinical forms of IBD are known: Crohn’s disease and ulcerative colitis. Even though the reason for IBD remains ambiguous, since several disease vulnerability genes have been recognized, genetic history is believed to be one of the reasons for IBD pathophysiology. However, genetic factors alone do not explain the quick rise in the prevalence of IBD, so environmental factors must also be important to its growth (Matsuoka and Kanai, 2015). In IBD patients, variations in the gut microbiome have been assessed that showed decreased gut diversity (Tong et al., 2013; Willing et al., 2010). Reduction in Firmicutes and increase in Proteobacteria are the most reliable findings of the modified composition of gut microflora in patients with IBD (Scanlan et al., 2006; Stecher and Hardt, 2008). The decline in the diversity of gut microflora has been observed due to the reduction in Firmicutes diversity in IBD patients. A decline in Clostridium leptum groups, in particular Faecalibacterium prausnitzii, among Firmicutes was recorded in a number of findings (Wang et al., 2014). Findings associated with species Enterobacteriaceae, Bacteroides, Bifidobacteria species, Lactobacillus species, and Escherichia coli are inconsistent with studies discussed above (Andoh et al., 2011; Takaishi et al., 2008).
There is a clear agreement on the involvement of gut microflora in the development of IBD, but the gut microflora alterations and involvement of specific bacterial species are under discussion. Prospective studies must be undertaken to investigate an evidence-based reason and the exact relationship between gut microflora and IBD. Such studies will have to be supported by experiments involving colonization of wild-type, germ-free, and genetically modified mice with individual bacterial species or a combination of various bacteria to identify the cause of bacterial strain and clarify the fate of the gut microflora in IBD.
Gut microflora in neurological disorders
Gut-brain axis terminology is used for biochemical signaling between the central nervous system (CNS) and the GI tract. A definition by Wang and Kasper describes the gut-brain axis that comprises the CNS, the neuroimmune and neuroendocrine systems, the parasympathetic and sympathetic sections of the autonomic nervous system, and gut microflora (Wang and Kasper, 2014). Research studies on the gut-brain axis have revealed the complex communication and interaction system for the proper maintenance of GI homeostasis. The efferent signals are derived by the autonomic system from the CNS to the intestinal walls (Arneth, 2018). Several studies indicate that through the communication and stimulation of “pattern recognition receptors” for example, toll-like receptors 2 and 4, the gut microflora influences the growth, functions, and ailments of the CNS and Enteric Nervous System (ENS) (Heiss and Olofsson, 2019; Hyland and Cryan, 2016). Gut dysbiosis and consequent loss of gut barrier stability and intestinal permeability enable improved translocation of gut-bacteria-derived metabolites and microbe-associated molecular patterns into mesenteric lymphoid tissues, leading to the progression and development of various neurological diseases (Tyler Patterson and Grandhi, 2020; Tremlett et al., 2017). An animal study also documented that fluctuations in the composition of intestinal microflora or lack of enteric bacteria in mice had a minor prevalence of myenteric neurons and a sophisticated incidence of intestinal motor dysfunction, suggesting that enteric bacteria had a determinable effect on ENS tropism (McVey Neufeld et al., 2013). In contrast to control mice, germ-free animals also displayed deregulated hormone signaling, less brain-derived neurotropic factor expression, neurotransmission variations, and amino acid metabolism (Kawase et al., 2017). Gut microbes modified the movement of locomotors in Drosophila by enhancing metabolite assembly (Chen et al., 2019). Researchers reported that various pathogenic, commensal, and probiotic microbes of the GI tract could activate the signaling process of CNS and neural pathways, and these can contribute to the development of depression and anxiety (Foster and Neufeld, 2013; Naseribafrouei et al., 2014). When studying the directional interactions between CNS and the gut, other agents include Lactobacillus rhamnosus, Helicobacter pylori, L. paracasei, Pseudomonas, Escherichia coli, and Bifidobacterium longum (Arneth, 2018). Recent research has focused on the contribution of the gut microflora to regulate and influence metabolism and immunity and how it can change the function of the brain. Other factors, including diet, sleep patterns, and exposure to antimicrobial agents, have been found to alter brain function by altering the gut microbiome (Arneth, 2018; Bravo et al., 2012).
Medical and scientific communities have increased awareness regarding the significant link between CNS and the intestinal environment. Specifically, existing research studies have shown that the gut-brain axis integrates the persistent interaction and bidirectional communication between the gut, the CNS, the endocrine system, the ENS, and the brain.
Gut microflora in cardiovascular diseases
Atherosclerosis is termed an inflammatory disorder with evidence supporting a potential autoimmune background (Hansson and Jonasson, 2009). Infection contributes to inflammation in the body and is considered a proposed mechanism of atherosclerosis. Various microbes are associated with an increased risk of cardiovascular diseases (CVD), including Chlamydophila pneumoniae, Helicobacter pylori, Porphyromonas gingivalis, influenza A virus, hepatitis C virus, cytomegalovirus, and human immunodeficiency virus (Rosenfeld and Campbell, 2011). Two predominant mechanisms are involved in the contribution of infections towards atherosclerosis: direct infection of the blood vessel wall that makes it susceptible to plaque formation or indirect infection at another site that promotes the proinflammatory mediators from a systemic immune response that can affect the growth of plaque (Novakovic et al., 2020). Gut dysbiosis involves atherosclerotic metabolite production in the gut, like trimethylamine N-oxide (TMAO), and it can change bile acid metabolism (Bu and Wang, 2018). The finding that DNA is found in atherosclerotic lesions from different species of bacteria and in the gut of similar persons indicates that the gut microflora can be a possible cause of atherosclerotic bacteria and consequently be involved in the coronary artery disease pathogenesis (Jin et al., 2019). Jie et al. (2017) studied the gut microflora and atherosclerotic CVD connection (Jie et al., 2017). They found that the gut microbiome composition involving Enterobacteriaceae and Streptococcus spp. members in atherosclerotic CVD was higher than that in healthy controls (Jie et al., 2017). Karlsson et al. (2012) used gut metagenome shotgun sequencing to show that the intestinal microbial populations differed from those in healthy controls in patients with symptomatic atherosclerosis (Karlsson et al., 2012). Patients had increased numbers of the genus Collinsella, while gender and age-matched controls had improved the richness of Eubacterium and Roseburia (Karlsson et al., 2012). Other evidence also exhibited the role of gut microflora in atherosclerosis in humans (Lanter et al., 2014).
To understand the role of gut microflora in human health and to guide the therapeutic interventions for CVD, it is important to elucidate the factors that work together to affect gut microflora and disease development. Further investigations are needed to examine these complex mechanisms by advanced nanomedicine approach, data sciences, and incorporation of various factors like ethnicity and sex to study the gut bacteria-mediated mechanisms that can lead to more effective preventive and therapeutic approaches to CVD.
Ancillary Effects of Inorganic Nanoparticles on Gut Microflora
Antibiotics widely used to treat bacterial infections can result in the imbalance of intestinal microflora, destroying the intestinal barrier and increasing bacterial resistance. There is an urgent requirement for a therapy that does not affect the intestinal microflora. NPs-based approaches can combat bacterial infections directly or indirectly to overcome antibiotic resistance without affecting the normal gut microflora. The potential effects of NPs on the microbiome and their clinical implications are still constrained in the number of studies. Recent evidence indicates that a number of NPs, including carbon nanotubes, titanium dioxides, cerium dioxides, cerium oxides, nano silica, and nano silver (Ag), may have an impact on microbiota (Lamas et al., 2020). Other NPs, such as the iron-NPs, may be more beneficial compared with standard iron-based additional drugs because they do not disturb the microbiota/microbiome (Pietroiusti et al., 2016). The treatment of intestinal diseases may also require NPs-based clinical treatments as they are reported as a safe mode of treatment as compared to antibiotics or other commercially available drugs (Qiu et al., 2018). Previous studies have revealed that Ag and copper (Cu) NPs were being used for antimicrobial action and in food packaging (Cushen et al., 2014), titanium dioxide as a food dye (Weir et al., 2012), and amorphous silica, introduced through the oral route, has been used as an alcoholic drink, clearing additives or as an anti-caking agent (Weir et al., 2012).
The pathophysiological pathways, biomarkers, and different metabolic activities in vitro and in vivo models following the interaction of gut microbiota with exposure to NPs have been discussed here. The potential toxicological effects NPs in various physiological and chemical processes induced by the microbiota are also highlighted. Fig. 2 represents the impact of inorganic NPs on gut microflora.
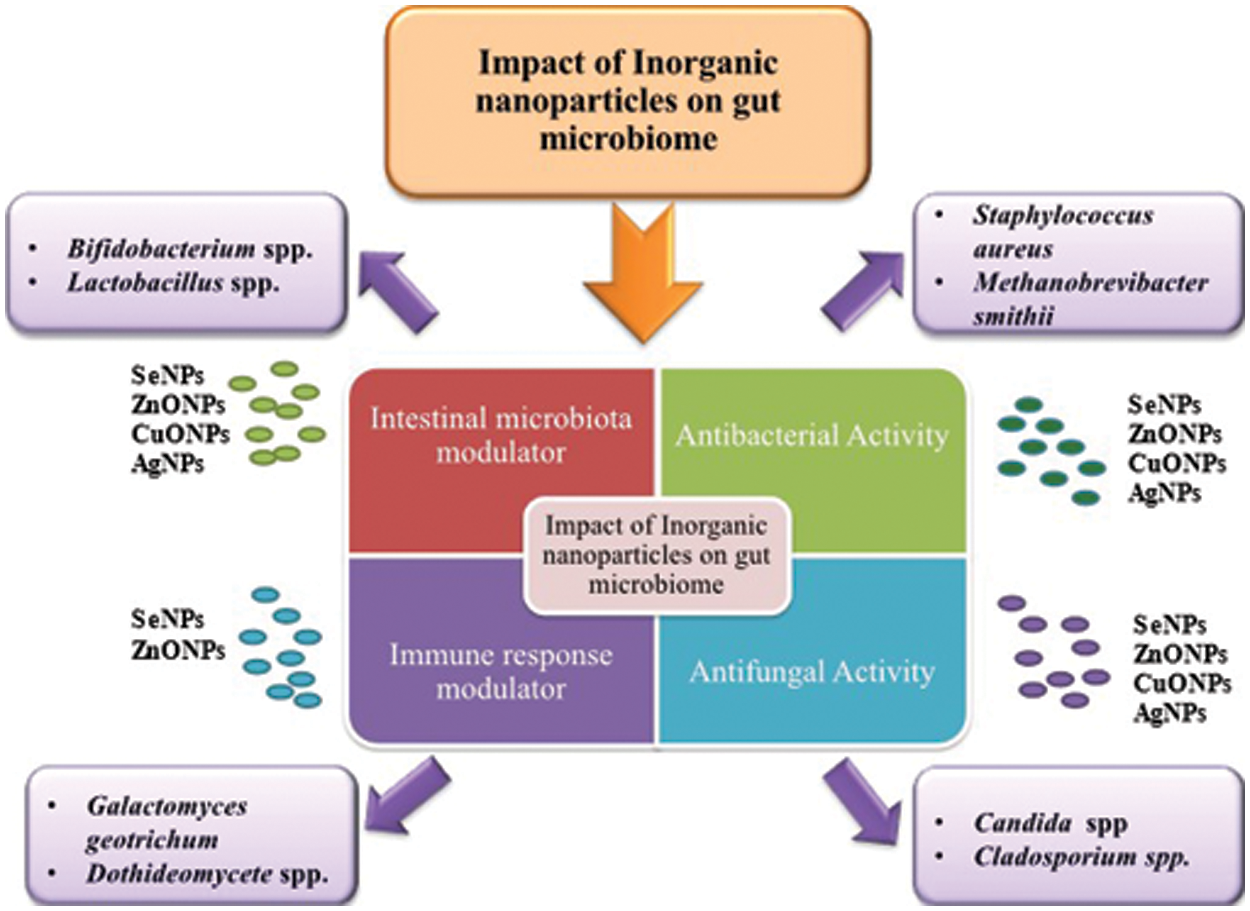
Figure 2: Role of inorganic nanoparticles (Se NPs, ZnO NPs, CuO NPs, Ag NPs) on gut microbiome as intestinal microbiota modulator, immune response modulator, antibacterial and antifungal agents.
Impact of Inorganic Nanoparticles on Gut Microflora of Animal Models
Se is considered a micronutrient used as a routine constituent of animal feed to maximally improve the efficiency of the immune system. Since then, mounting studies have shown the beneficial effect of the organic and inorganic Se supplementation on immune-endocrine, metabolic, and cell homeostasis of Se (Wang et al., 2017). The inorganic (selenite) and organic (selenomethionine) formulations are currently delivered in two separate forms (Dekkers et al., 2013). Recently, the potential application of this micronutrient in the form of Se NPs (Se-NPs) has been emphasized (Surai, 2002a, 2002b), and it has also been shown to be responsible for modulating gut microbiota (Gangadoo et al., 2016). Previous studies revealed that administration of Se in the form of NPs increased the uptake of Se by the cells, which led to improved immune responses and modulation of intestinal microbiome and musculoskeletal functions (Beski et al., 2015; Gangadoo et al., 2018; Lin et al., 2015). Due to the best absorption ability of NPs, SeNPs are considered the most appealing material that does not go through the metabolism before being incorporated into selenoproteins as their absorption mechanism is much faster because of their unique physical property of high surface-to-volume ratio (Surai, 2002b; Wang et al., 2011a).
Zn is the second most widely dispersed micronutrient found in traces (Ross et al., 2020; Faizan et al., 2021). It is present in a variety of foods, including beef, oxygen, meals, and grains. Commercial Zn supplements have between 7 and 80 mg of elementary Zn and are usually formulated as zinc oxides or acetate-gluic salts (Ross et al., 2020). Commercial Zn supplements contain between 7 and 80 mg of primary Zn and are typically formulated as genetically engineered Zn supplements. Symptoms of zinc deficiency are nonspecific, including growth retardation, diarrhea, alopecia, glossitis, nail dystrophy, decreased immunity, and hypogonadism in males. In developing countries, Zn supplementation may be effective for the prevention of upper respiratory infection and diarrhea and as an adjunct treatment for diarrhea in malnourished children (Mocchegiani et al., 1995; Timbo et al., 2006). ZnO NPs are considered a rich source of Zn and are less than 100 nm in diameter. For many years it has been used as an anti-cancer, antibacterial, antifungal, anti-viral, and in industrial products (Karavolos and Holban, 2016; Patel et al., 2016). Under several circumstances, the detrimental effects of ZnO NPs on animals have been observed, and these harmful effects are closely associated with NP size and form. More toxic effects arise because NPs of the smallest scale quickly pass across the cell membrane, thus averting the defense mechanism (Handy et al., 2008). Then, NPs migrate into the cell to enter mitochondria, alter the metabolism of the cell and contribute to cell death (Saptarshi et al., 2015). ZnO NPs are responsible for the antibacterial activity as the reduced particle size induces surface reactivity (Crisol-Martínez et al., 2017). The reason behind the antibacterial effect is the production of reactive oxygen species (ROS), which is responsible for the induction of oxidative stress in the bacterial cell (Crisol-Martínez et al., 2017).
In an experimental study to examine the effect of a low dose of ZnO NPs performed on weaned piglets, ZnO NPs in the diet increased the amount and diversity of bacteria in the ileum and decreased both in cecum and colon, while the relative abundances of Streptococcus and Lactobacillus increased, respectively. In addition, mouse urine metabolites (Yan et al., 2012) and impaired glucose metabolism in lung epithelial cells suggested the effect of ZnO NPs on metabolism (Lai et al., 2015).
Furthermore, the impact of ZnO NPs on the intestinal microflora of domestic animals and the association between the metabolites and the intestinal microflora is not yet known. Further research work is needed to find the beneficial or harmful effects of ZnO NPs on the bacterial population of ileal digesta, the metabolites present in the plasma, and the inter-relationship because intestinal microflora is of great importance to metabolism (Sirelkhatim et al., 2015).
Cu is an important micronutrient for the functioning of living cells and can be found in both oxidized and reduced forms. Furthermore, various enzymes involved in metabolic processes, specifically to the proper functioning of the body, are built into the active centers (Fröhlich and Fröhlich, 2016; Xia et al., 2017), such as mitochondrial respiration, protection against free radicals, neurotransmitter synthesis, development of collagen and elastin, synthesis of melanin, or metabolism of iron (de Bie et al., 2007; Festa and Thiele, 2011; Lutsenko et al., 2007). It has been reported that excessive use of Cu has major toxic effects that will lead to extreme metabolic syndromes such as Menkes disease or Wilson’s disease (Gupta and Lutsenko, 2009; Vickers, 2017). For instance, CuO nanoparticles (CuO NPs) are used in conductive coatings, batteries, surfactants, and antimicrobials (Tümer and Møller, 2010). In general, because of their high surface-to-volume ratio, CuO NPs can exhibit efficient antimicrobial activity in a wide variety of microbial species at low concentrations (Hyland and Cryan, 2016). Cu-based NPs are also increasingly being used commercially, and their worldwide production in 2010 reached 200 tons (Lorincz, 2018).
Recently, a study has been performed on Cyprinus carpio (common carp) to assess the effect on growth, immunity, and oxidation resistance of common carp (3.02 ± 0.01 g, original average weight ± S.E.) of Cu NPs (Dawood et al., 2020). Five fish classes were fed with 0, 0.5, 1, 2, and 4 mg/kg inorganic Cu diets for eight weeks. The growth rate of Cu-NPs in diets improved, and the feed conversion ratio decreased with the linear and quadratic model (P < 0.05). Increased protein, lipid, and ash content were also found to be dose-dependent in popular carp Cu-NP (P < 0.05). Cu deposition improved by Cu-NPs in carcass, liver, muscle, and gills, with an excess of 4 mg Cu-NPs/kg (P < 0.05) (Dawood et al., 2020). Cu deposition improved by Cu-NPs in carcass, liver, muscle, and gills, with an excess of 4 mg Cu-NPs/kg (P < 0.05). Blood variables with the exception of HB, RBC, total proteins, albumin, and globulins with the maximum amounts of 2 mg/kg (P < 0.05), have not been substantially altered by supplementation with Cu-NP. The Cu-NPs with reduced malondialdehyde content (P < 0.05) enhanced the amount of IgM, phagocytic, lysozyme, SOD, CAT, or GPX activity. Based on the regression analysis, the requirement of common carp dietary Cu NPs has been estimated to be 2.19 to 2.91 mg/kg (Dawood et al., 2020). Another study reported the in vivo exposure impacts of biocidal nanomaterial exposure on intestinal microbiota, host immune responses, and the host susceptibility to earthworm bacterial disorders. Eisenia fetida was exposed to CuO NPs in soil for 28 days, after which the soil bacterium Bacillus subtilis threatened the existence of earthworms. Immune responses to identify earthworm immune genes were calculated by measuring the mRNA levels (Swart et al., 2020). Treatment implications for the intestinal microbiota have also been tested for associating changes in the microbiome with immune reactions. Treatments have indeed been responsible for a change in the gut microbiota of earthworms. No effects of therapy on earthworm immune marker expression were reported despite these effects (Swart et al., 2020).
Recently it has been reported that Ag NPs are the major constituents of consumer goods, and are used as food additives and food contact materials, mainly due to their antimicrobial properties (Jarosz et al., 2018; Scott et al., 2018b). Ag NPs have achieved a revived, rising interest as an antimicrobial substitute that could substitute or supplement the activity of food chemical preservatives (Chaudhry et al., 2008; El-Katcha et al., 2020). In this context, the possible application of two Ag NPs, PEG-AgNPs 20 and glutathione (GSH)-AgNPs, have been observed to regulate microbial processes in winemaking (Chaudhry et al., 2008). To promote their environmental interactions, Ag NPs should be coated; a GSH coating enhances the solubility and ability of Ag NPs to interact with the environment.
GSH-AgNPs have been reported as potential antimicrobial agents against the foodborne pathogen Campylobacter and were also utilized in the manufacturing and processing of meat in the poultry industry (Monge and Moreno-Arribas, 2016), with an average Ag dietary intake of 70–90 μg per day, or even higher (García-Ruiz et al., 2015; Gil-Sánchez et al., 2018). Another study was performed to investigate the occurrence of morphological and physicochemical changes in the GI tract after the ingestion of Ag NPs. Ag NPs get modified in size and morphology when exposed to saliva, gastric juice, and intestinal fluids. These changes are due to differences in pH and composition of each fluid (Silvan et al., 2018). Besides, these NPs are less toxic and can also act as anti-inflammatory agents (Sergeevna et al., 2018). Another study was performed by using a stable and coherent model framework to verify the complex movement of fluids across all simgi® (a computer-controlled GI in vitro model design) compartments (Wijnhoven et al., 2009). This study also involved the development of the full function of the simgi® dynamic GI simulator, which has proved to be a valuable instrument for determining the physiological behavior of Ag NPs. In the transport of Ag along the simgi®, a similar dilution pattern was also found for both volunteers, supporting the model’s reproducibility. Ag NPs have undergone several GI fluid transformations. Overall, during the complex GI simulations of Ag NPs at simgi®, there were no improvements in bacterial composition or production of ammonium ions. This seems to suggest that the structure and metabolic function of human intestinal microbiota was not disrupted by these nanomaterials, which is of considerable importance in view of its possible use in the area of the food industry (Walczak et al., 2012).
In animal nutrition, Ag NPs can be used as prebiotics (Ognik et al., 2016). A research study on weaned piglets by Fondevila et al. (2009) showed the administration of 20 and 40 ppm of Ag NPs with reduced coliform bacteria of ileal contents. Pathogenic bacteria, Clostridium perfringens/Clostridium histolyticum group, was considerably decreased by Ag NPs with 20 ppm concentration, but the other major ileum bacterial groups were not impacted (Ognik et al., 2016). A study on mice by Wilding et al. (2016) did not change the gut microflora of mice after a repeated dose of Ag NPs (20 or 110 nm) administered for 28 days with either citrate coatings or PVP (Wilding et al., 2016). In this study, the Ag NPs dose corresponded to 2000× the oral reference dose; the findings show that for colloidal Ag, daily intake was considered safe over a lifetime in humans (Bergin and Witzmann, 2013). Some contradictory studies also showed the toxic effects of Ag NPs after their ingestion at higher levels. Further long-term chronic toxicity studies are needed.
According to some studies, TiO2 NPs have low distinct effects on the microbiome. Toxicity studies on TiO2 NPs showed only a minor decrease in Bacteroides ovatus and an increase in Clostridium cocleatum, leading to the conclusion that TiO2 NPs at low concentrations have no major impact on the gut microflora (Ghebretatios et al., 2021). However, another study showed the exposure of mice to 2.5 mg/kg of TiO2 NPs for 7 days found no alterations in the fecal microbiota composition (Chen et al., 2017). Another study to check the impact of TiO2 NPs on gut microflora using an in-vitro Human Gut Simulator system found that community density was reduced, but no impact was observed on diversity, microbial functionality, and fermentation (Agans et al., 2019). These studies describe the little impact of TiO2 NPs on the gut microbiome.
By contrast, other studies showed that TiO2 NPs could alter the gut microflora. Daily oral administration of TiO2 NPs with 0, 2, 10, and 50 mg/kg doses to rats for 30 days could induce gut dysbiosis like an increase in Lactobacillus gasseri, Turicibacter, and Lactobacillus NK4A136_group and a reduction in Veillonella (Utembe et al., 2022).
SiO2 NPs have not been studied extensively in terms of their impact on the gut microbiome. However, one study showed the negative impact of SiO2 NPs on the gut microflora. Researchers found enhanced diversity and richness of the microbial community after exposure of mice to human-relevant doses of SiO2 NPs (2.5 mg/kg) for one week (Chen et al., 2017); Firmicutes and Proteobacteria increased, and Bacteroidetes and Lactobacillus were found to decrease. The absorption rate of precipitated or fumed silicate allows it to accumulate in the gut lumen, which provides more time for toxic effects on gut microflora. It is concluded that fumed silicates have more potential harm (Ghebretatios et al., 2021).
Impact of Inorganic Nanoparticles on Gut Microflora of Poultry
Inorganic NPs have various applications in the poultry industry, including rapid and specific diagnosis of disease, production parameters improvement in broilers, microbial inhibition, immuno-stimulation, and disinfection. It is important to well assess the safety and toxicity hazards before the application of NPs in poultry farms to guarantee the health of both poultry and humans.
Se is a micronutrient used in the poultry industry for the modulation of the gut microbiome. Several studies have already been used to monitor pathogenesis by using different vaccines and antibiotics. The increase in antibiotic resistance, however, has led researchers, as well as chicken farmers, to pursue alternate methods to deal with animal and human pathogens (Obeng et al., 2012; Suzuki and Ogra, 2002). New and emerging methods of nanotechnology have been found not only to kill pathogenic bacteria but also to resolve toxicity and consequent bioaccumulation (Vieira de Souza et al., 2012). The effect of Se NPs on pathogen production control was investigated in the poultry industry to improve the productivity, health, and wellbeing of the flocks. The experimental design of the poultry flock study was designed to compare the effect of bulk Se macronutrients used in the poultry industry with that of Se NPs. Se NPs, at 0.9 mg/kg concentrations, have demonstrated the highest productivity by increasing the abundance of beneficial bacteria such as Lactobacillus and Faecalibacterium (Gangadoo et al., 2016). It also increases the concentration of SCFA, including butyric acid, to boost intestinal health. Butyric acid is considered a major intestinal metabolite used to store energy for colonic cells and other significant bodily functions (Gangadoo et al., 2016).
In chickens, the richness and variety of cecum microbiota were altered with chronic deficiency of dietary Zn (Reed et al., 2015). SCFA and monoglycerides have been reported as antibacterial agents against Campylobacter jejuni infection in broiler poultry (Guyard-Nicodeme et al., 2016). It was observed that Zn-regulated cecal microbial population by increasing the number of beneficial bacterial species like Lactobacillus and decreasing the number of Salmonella in broilers. Relevant improvements in the cecal microbiota of broiler chicken were caused by the antibiotic zinc-bacitracin (Costa et al., 2017; Gong et al., 2007; Shao et al., 2014; Torok et al., 2011).
Several studies have compared different concentrations and states of Cu (organic, inorganic, and nano) to evaluate the effect on growth enhancement, immune response, and blood biochemistry in poultry (Du et al., 2019; Ho et al., 2018; Scott et al., 2018a; Zhang et al., 2014). A previous study on broiler chickens investigated the effects of dietary substitution of inorganic Cu on growth efficiency, immune response, intestinal microbiota, and intestinal microbiota using the same (100 percent of the recommended requirements) or lower (50 percent of the recommended requirements) levels as organic Cu or Cu-NPs with a source of fresh or oxidized oil (El-Kassas et al., 2018). In the assessment of 50% organic Cu and Cu NPs, various activities were increased and it revealed bactericidal activity, phagocytosis, and lysosomal events, while high doses reduced these activities. On the other hand, organic Cu and Cu-NPs decreased the number of Escherichia coli and Lactobacilli in the cecum of chicken but significantly decreased the inflammation rate of liver tissues, proving it as an anti-inflammatory agent (El-Kassas et al., 2018). Fig. 3 shows the impact of NPs on the poultry.
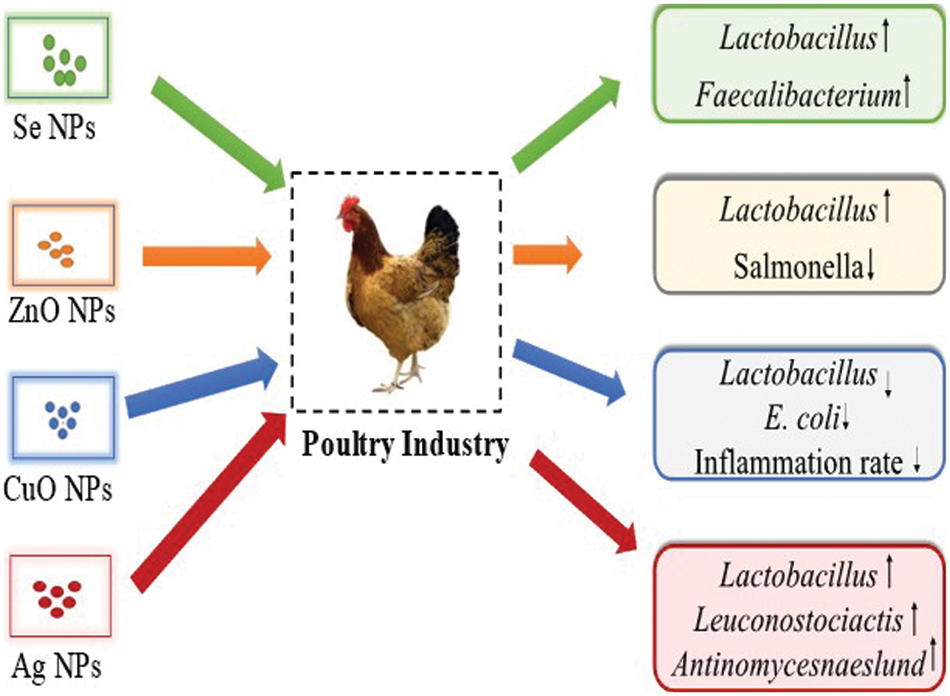
Figure 3: The impact of various inorganic nanoparticles on poultry by increasing the commensal bacteria and reducing the pathogenic microbes.
Ag NPs are considered an important feed supplement in the poultry industry. There is very little data on the impact of Ag NPs on gut microbes, and it needs further investigation as these NPs affect this ecosystem both positively and negatively. Sawosz et al. (2007) evaluated the effects of Ag NPs on the cecal microbial profile and enterocyte morphology in the Japanese quail duodenum. Ten days old poultry model quails were divided into four groups with 15 quails in each group and placed for 12 days into four cages. Ag NPs with varying concentrations of 0, 5, 15, and 25 mg/kg were administered to quails via drinking water, and their cecal and duodenum microflora were collected at the end of the experiment by killing them. Initially, AgNPs did not affect the quail cecal microflora emphatically, but at 25 mg/kg in water, there was a remarkable elevation in the lactic acid bacterial population. Ag NPs did not show any deleterious effects on enterocytes in the duodenal villi (Sawosz et al., 2007).
In another study, when Ag NPs were fed to broilers, there was a decrease in the abundance of pathogenic Escherichia coli as compared to the control group, and the beneficial bacteria Lactobacillus remained unaffected. Un-sexed seven-day-old broiler chicks “Hubbard” (total 180) were divided into six groups with each group of three replicates, and 10 birds in each replicate were supplemented with varying concentrations of Ag NPs at 2, 4, 6, 8, and 10 ppm/kg during the growth trial period for 7–35 days. Ag NPs at 4 ppm/kg led to the best productive performance of broilers (Elkloub et al., 2015).
Further research is needed in this area to investigate the effects of Ag NPs on beneficial bacteria of the gut in the poultry industry.
Response of Pathogenic Bacteria to Nanoparticles
To enhance their growth and virulence, pathogenic bacteria residing in the gut cells utilize the microbiota-derived nitrogen and carbon sources as nutrients and regulatory signals giving rise to recurrent infections (Campoy and Colombo, 2009). Enteric pathogenic bacteria include Citrobacter rodentium, Salmonella enterica serovar Typhimurium, Shigella flexneri, Clostridium difficile, and Listeria monocytogenes (Bäumler and Sperandio, 2016). Clostridium Difficile transmitted via the fecal-oral route is a Gram-positive toxin-producing bacterium. Release of two toxins from Clostridium Difficile as an enterotoxin causing enhanced intestinal permeability and secretion of fluid and a cytotoxin leading to a strong colonic inflammation resulting in colitis, diarrhea, and cell death (Ofosu, 2016). Expansion of Clostridium difficile population occurs due to the antibiotics treatment that reduces the microbiota diversity in the gut (Bäumler and Sperandio, 2016). Common facultative intracellular pathogens Salmonella enterica serovar Typhimurium, a primary enteric Gram-negative pathogen, infects millions of people each year and causes life-threatening foodborne infections (Eng et al., 2015; Fàbrega and Vila, 2013). GI diseases trigger when this bacterium reaches the intestinal epithelium. Antibiotic treatment elevates the levels of free sialic acid from the host and succinate from the microbiota in the intestinal lumen promoting the expansion of the S. typhimurium population and can cause gastroenteritis if it enters the intestinal epithelial cells (Bäumler and Sperandio, 2016). Survival and replication of Salmonella species in the host cells, including macrophages, have been reported. This intracellular bacterial localization complicates the antibiotic treatment due to their limited capability to cross the mammalian cell membranes and it can be exported out actively by the host cell (Kamaruzzaman et al., 2017). Due to the high drug loading capacity and their capability to penetrate the eukaryotic cells, NPs, on the other hand, can overcome this challenge. The strategies for treating infections caused by Enterobacter species are restricted due to the faster growth rate of resistance, against last-resort antibiotics. Nanomaterials are confined to no resistance development and can deliver a sustainable therapeutic design. Unlike antibiotics, NPs with access to multiple targets can possess multiple killing mechanisms and specific biological structures, enabling them to escape from enzyme deactivation, ultimately leading to decreased resistance (Pelgrift and Friedman, 2013). Endocytosis and membrane fusion as non-porin mechanisms used by nanomaterials enable their entry into bacterial cells (Pelgrift and Friedman, 2013). Efflux pumps can be blocked by nanomaterials to enhance the accumulation of antibiotics in the bacterial cell (Gupta et al., 2017). Nanomaterials are active against persisters because their killing mechanism due to membrane damage does not involve the bacteria being in the active growth state (Hurdle et al., 2011). NPs possess specific surface chemistry that allows easy penetration of nanomaterials into bacterial biofilms, rendering the interaction to deeply entrench bacterial cells. Significant interactions with extracellular polymeric substances are aided by the amphiphilic balance of many nanostructures, like hydrophobic and electrostatic interactions, optimizing adsorption, and diffusion through biofilms (Gupta et al., 2016).
Mechanism of nanoparticles against pathogenic bacteria
Nanomaterials with unique sizes and shapes can target pathogenic bacteria by various bactericidal mechanisms like cell wall and cell membrane damage, production of ROS, and finally binding to the intracellular components of the cell (Soenen et al., 2011). NPs possess exceptional physiochemical characteristics, specifically multivalent interactions like receptor-ligand interactions, electrostatic interactions, van der walls forces, and hydrophobic interactions with bacteria (Makabenta et al., 2021).
Disruption of the cell wall and cell membrane
The bacterial cell envelope has evolved to act as an antimicrobial physical barrier. Gram-positive bacterial cell walls possess teichoic acids, and Gram-negative bacterial cell walls possess lipopolysaccharides; teichoic acids possess phosphate groups rendering the bacterial surfaces negatively charged. Better electrostatic interactions of positively charged nanomaterials are established with negatively charged bacteria compared to mammalian cells, which are less negatively charged (Makabenta et al., 2021). To disrupt the bacterial membrane selectively, the hydrophobicity and charge density of NPs act as important factors in designing the nanomaterials (Huo et al., 2016; Makabenta et al., 2021). Selectivity decreases when NPs are overly hydrophobic, and with a high cationic charge, can bind to the mammalian cell surface. A strong antimicrobial effect with reduced hemolysis and cytotoxicity levels of cationic nanomaterials can be generated by optimizing the balance between cationic charge and hydrophobicity, creating a good amphiphilic balance (Makabenta et al., 2021).
Reactive oxygen species generation
Cell signaling, differentiation, survival, and death are highly affected by cellular oxidative metabolic processes by-products known as ROS. Through the excessive accumulation of ROS, lethal oxidative stress occurs to cause damage by various mechanisms, especially superoxide and hydroxyl radicals reaction to protein thiols and deactivating membrane receptors (Memar et al., 2018). NPs generate ROS species by directly producing from the surface of NPs or their leached ions, by interacting with intracellular organelles, and oxidation by interacting with redox-active biomolecules like NADPH oxidase (Miller et al., 2015). Due to the intrinsic photocatalytic activity of NPs, some metal NPs utilize ROS production as the main antibacterial mechanism (Memar et al., 2018; Miller et al., 2015).
Damage to the intracellular components
For bacterial cell functioning and survival, cellular homeostasis and intracellular signaling pathways have the core importance. So, the NPs intrude these processes, eventually leading to the death of bacterial cells. NPs can disturb and alter gene expression, DNA damage, mitochondrial damage, and synthesis of proteins (Shamaila et al., 2016). Fig. 4 represents the antibacterial mechanism of action of inorganic NPs.
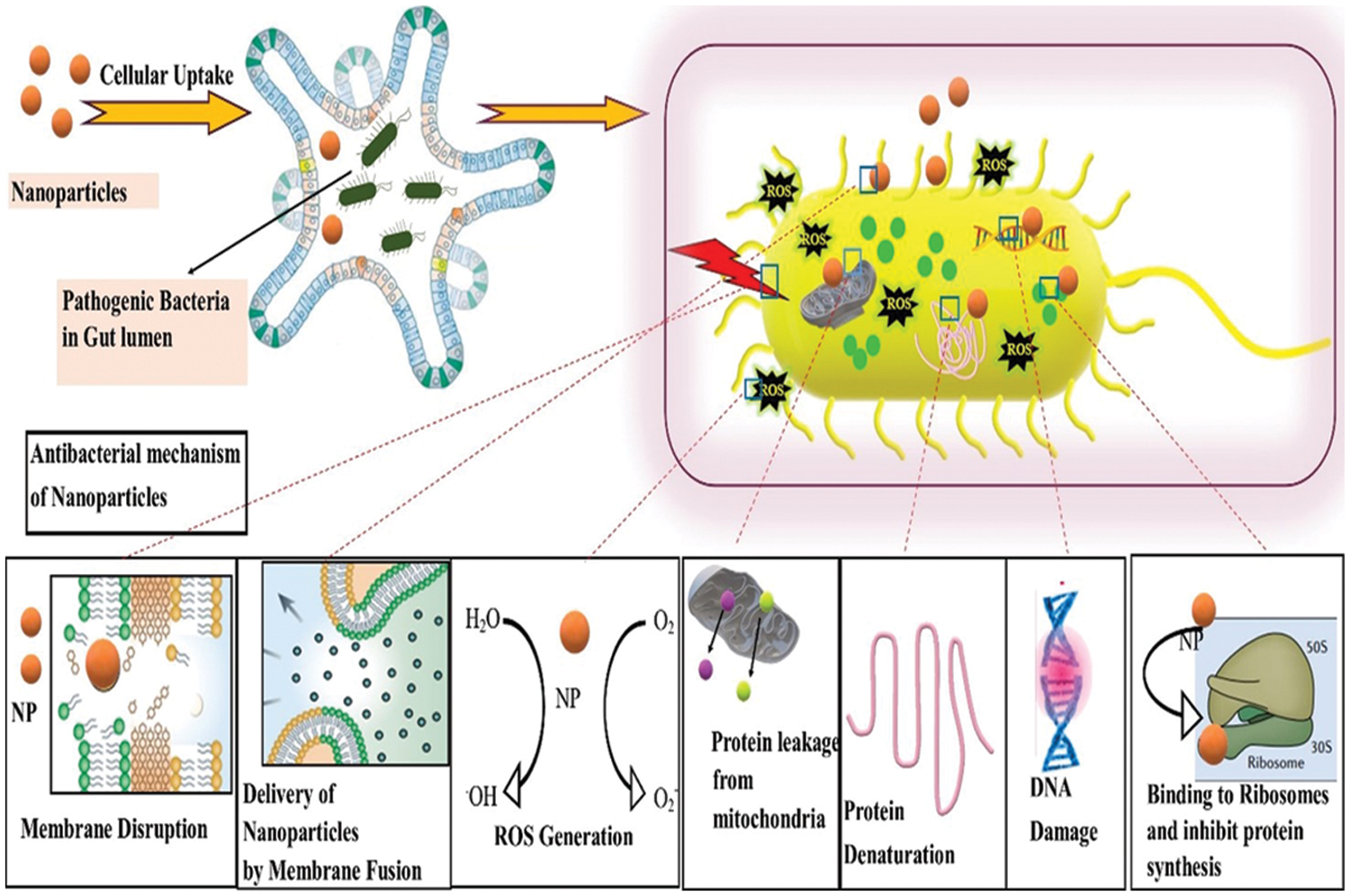
Figure 4: Antibacterial mechanism of inorganic nanoparticles (NPs). The uptake of NPs to the gut lumen eventually affects the pathogenic bacteria residing inside the cell. Bacterial diameter is approximately 0.2–10 µm, and that of NPs ranges from 1–100 nm, rendering its strong interaction to bacterial membrane and utilizing a variety of bactericidal mechanisms.
Generally, nanomaterials ideally formulated with unique sizes, shapes, and physiochemical properties may provide several bactericidal pathways to tackle bacteria that evade antibiotic resistance mechanisms and offer a broad design space for novel antimicrobial agents.
Opportunities, Challenges and Future Perspectives
For the treatment of recalcitrant multidrug-resistant bacterial infections, NPs offer an evolving ‘outside of the box’ strategy. The configurable properties of nanomaterials, especially their surface functionality, offer design spaces that can quite well optimize the therapeutic effect but reduce the host toxicity. Inorganic NPs interfere with normal GIT flora when present in GI fluids. Small size with a greater surface area of NPs provides a larger area for absorption of any surface-active elements in the GIT. Subsequently, the rate of digestion of lipids, protein, or starch could reduce by higher NPs levels in GIT. Aggregation of inorganic NPs in the GIT can decrease their exposed surface area. Very little data are available for the potentially harmful impacts the NPs in the GIT. Starch, lipid, and protein can be digested fully by additional enzymes and other digestive components secreted by the body. Normally, a very low level of NPs is ingested, so this mechanism is not considered a major health concern (McClements and Xiao, 2017).
Some inorganic NPs may disrupt the crucial structures in the GIT, like microvilli, and change the normal functioning of epithelial cells and normal nutrient absorption (Fröhlich and Fröhlich, 2016). Cytotoxicity of inorganic NPs could result from multiple mechanisms, but the most important is the production of ROS species that may damage the cell membrane, cell organelles, and nucleus when interacting with proteins, lipids, and nucleic acids (Wu et al., 2014). However, it is still unknown to which extent inorganic NPs could produce cytotoxicity during their consumption as a complex diet in normal conditions. NPs can interact with the beneficial bacteria in the gut and could change their viability to possibly adversely affect health as some bacterial species have an important role in human health (McClements and Xiao, 2017). He determination of NPs’ impact on beneficial gut microbiota is an important research area.
The review by (McClements and Xiao, 2017) suggests that some types of NPs are cytotoxic while some others are not so; the reasons for these inconsistencies are variable physiochemical properties of NPs, including internal and surface composition, crystal form, physical state, shape, dimensions, aggregation state, crystal form, and dose. In some research studies, NPs properties have not been sufficiently characterized, and variable testing methods are used to determine their possible toxicity, including physiochemical, cell culture methods, microbial, animal, and human studies. Furthermore, test methods tend to change from lab to lab, thus complicating a direct comparison of results. Major impacts of these factors on NPs properties, their behavior, and toxicity need to be studied (McClements and Xiao, 2017).
The above discussion clearly indicates that novel standardized strategies need to be developed to effectively test the toxicity of inorganic NPs in realistic and reproducible conditions. In the future, NPs can be used as a vehicle for delivery, absorption, and bioavailability of polyphenols on gut microbiota, as reviewed by Cardona et al. (2013), who elaborated in detail the role of polyphenols in gut microbiota. Intake of polyphenols could reduce the pathogenic bacterial population and increase the commensal microbes like Bifidobacterium and Lactobacillus and exert a prebiotic-like effect (Cardona et al., 2013).
Human biology is concerned with concomitant micro-organisms, most of which reside in the digestive tract, from which different chemicals are released or modified or cause host reactions that influence different physiological functions, including immunity, neurobiology, and metabolism. The shift in the co-existence of microbial genera contributes to dysbiosis, resulting in many human disorders. Diet and diet-related agents may possess a direct impact on the health of the host by controlling the gut microbiome, which can thus sustain the gut’s homeostasis. Analysis of the GI microbiota and agents that can manipulate the intestine requires a thorough understanding. For decades, nanotechnology has been utilized as a tunable platform that could be adapted for unique obstacles and challenges. Overall, the use of nanotechnology in microbiome modulation is still a nascent field, but these studies illustrate the potential at this growing intersection. The production of suitable in vitro and in vivo models demonstrating the effectiveness and protection of NPs would ensure the clinical viability of their usage. This review, therefore, focuses on the effects on the gut microbiota and its positive, or negative influence of metal-based nanoparticles. It has been concluded that in the coming future, nanotechnology will replace the commercially available drugs and medicines used for the treatment of gut dysbiosis.
Author Contributions: SA and MAB jointly wrote this review. SA drafted the manuscript. BU, SA and MAB contributed to the in-depth discussion and conception. BAK and FM revise and finalize the manuscript. All authors approved the final version of the manuscript.
Ethics Approval: Not applicable.
Funding Statement: The authors received no specific funding for this study.
Conflicts of Interest: The authors declare that they have no conflicts of interest to report regarding the present study.
References
Agans RT, Gordon A, Hussain S, Paliy O (2019). Titanium dioxide nanoparticles elicit lower direct inhibitory effect on human gut microbiota than silver nanoparticles. Toxicological Sciences 172: 411–416. DOI 10.1093/toxsci/kfz183. [Google Scholar] [CrossRef]
Andoh A, Imaeda H, Aomatsu T, Inatomi O, Bamba S, Sasaki M, Saito Y, Tsujikawa T, Fujiyama Y (2011). Comparison of the fecal microbiota profiles between ulcerative colitis and Crohn’s disease using terminal restriction fragment length polymorphism analysis. Journal of Gastroenterology 46: 479–486. DOI 10.1007/s00535-010-0368-4. [Google Scholar] [CrossRef]
Angelakis E, Armougom F, Million M, Raoult D (2012). The relationship between gut microbiota and weight gain in humans. Future Microbiology 7: 91–109. DOI 10.2217/fmb.11.142. [Google Scholar] [CrossRef]
Arneth BM (2018). Gut-brain axis biochemical signalling from the gastrointestinal tract to the central nervous system: Gut dysbiosis and altered brain function. Postgraduate Medical Journal 94: 446–452. DOI 10.1136/postgradmedj-2017-135424. [Google Scholar] [CrossRef]
Ba Q, Li M, Chen P, Huang C, Duan X, Lu L, Li J, Chu R, Xie D, Song H (2017). Sex-dependent effects of cadmium exposure in early life on gut microbiota and fat accumulation in mice. Environmental Health Perspectives 125: 437–446. DOI 10.1289/EHP360. [Google Scholar] [CrossRef]
Bäumler AJ, Sperandio V (2016). Interactions between the microbiota and pathogenic bacteria in the gut. Nature 535: 85–93. DOI 10.1038/nature18849. [Google Scholar] [CrossRef]
Bergin IL, Witzmann FA (2013). Nanoparticle toxicity by the gastrointestinalroute: Evidence and knowledge gaps. International Journal of Biomedical Nanoscience and Nanotechnology 3: 1–2. DOI 10.1504%2FIJBNN.2013.054515. [Google Scholar]
Beski SS, Swick RA, Iji PA (2015). Specialized protein products in broiler chicken nutrition: A review. Animal Nutrition 1: 47–53. DOI 10.1016/j.aninu.2015.05.005. [Google Scholar] [CrossRef]
Bist P, Choudhary S (2022). Impact of heavy metal toxicity on the gut microbiota and its relationship with metabolites and future probiotics strategy: A review. Biological Trace Element Research 200: 5328–5350. DOI 10.1007/s12011-021-03092-4. [Google Scholar] [CrossRef]
Bravo JA, Julio-Pieper M, Forsythe P, Kunze W, Dinan TG, Bienenstock J, Cryan JF (2012). Communication between gastrointestinal bacteria and the nervous system. Current Opinion in Pharmacology 12: 667–672. DOI 10.1016/j.coph.2012.09.010. [Google Scholar] [CrossRef]
Bu J, Wang Z (2018). Cross-talk between gut microbiota and heart via the routes of metabolite and immunity. Gastroenterology Research Practice 2018: 1–8. DOI 10.1155/2018/6458094. [Google Scholar] [CrossRef]
Buttó LF, Haller D (2016). Dysbiosis in intestinal inflammation: Cause or consequence. International Journal of Medical Microbiology 306: 302–309. DOI 10.1016/j.ijmm.2016.02.010. [Google Scholar] [CrossRef]
Campoy E, Colombo MI (2009). Autophagy in intracellular bacterial infection. Biochimica et Biophysica Acta (BBA)-Molecular Cell Research 1793: 1465–1477. DOI 10.1016/j.bbamcr.2009.03.003. [Google Scholar] [CrossRef]
Cardona F, Andrés-Lacueva C, Tulipani S, Tinahones FJ, Queipo-Ortuño MI (2013). Benefits of polyphenols on gut microbiota and implications in human health. The Journal of Nutritional Biochemistry 24: 1415–1422. DOI 10.1016/j.jnutbio.2013.05.001. [Google Scholar] [CrossRef]
Chaudhry Q, Scotter M, Blackburn J, Ross B, Boxall A, Castle L, Aitken R, Watkins R (2008). Applications and implications of nanotechnologies for the food sector. Food Additives and Contaminants 25: 241–258. DOI 10.1080/02652030701744538. [Google Scholar] [CrossRef]
Chen H, Zhao R, Wang B, Cai C, Zheng L, Wang H, Wang M, Ouyang H, Zhou X, Chai Z (2017). The effects of orally administered Ag, TiO2 and SiO2 nanoparticles on gut microbiota composition and colitis induction in mice. NanoImpact 8: 80–88. DOI 10.1016/j.impact.2017.07.005. [Google Scholar] [CrossRef]
Chen K, Luan X, Liu Q, Wang J, Chang X, Snijders AM, Mao JH, Secombe J, Dan Z, Chen JH (2019). Drosophila histone demethylase KDM5 regulates social behavior through immune control and gut microbiota maintenance. Cell Host & Microbe 25: 537–552, e538. DOI 10.1016/j.chom.2019.02.003. [Google Scholar] [CrossRef]
Clemente JC, Pehrsson EC, Blaser MJ, Sandhu K, Gao Z, Wang B, Magris M, Hidalgo G, Contreras M, Noya-Alarcón Ó (2015). The microbiome of uncontacted Amerindians. Science Advances 1: e1500183. DOI 10.1126/sciadv.1500183. [Google Scholar] [CrossRef]
Costa MC, Bessegatto JA, Alfieri AA, Weese JS, Filho JA, Oba A (2017). Different antibiotic growth promoters induce specific changes in the cecal microbiota membership of broiler chicken. PLoS One 12: e0171642. DOI 10.1371/journal.pone.0171642. [Google Scholar] [CrossRef]
Cox LM, Blaser MJ (2015). Antibiotics in early life and obesity. Nature Reviews Endocrinology 11: 182–190. DOI 10.1038/nrendo.2014.210. [Google Scholar] [CrossRef]
Crisol-Martínez E, Stanley D, Geier MS, Hughes RJ, Moore RJ (2017). Understanding the mechanisms of zinc bacitracin and avilamycin on animal production: Linking gut microbiota and growth performance in chickens. Applied Microbiology and Biotechnology 101: 4547–4559. DOI 10.1007/s00253-017-8193-9. [Google Scholar] [CrossRef]
Cushen M, Kerry J, Morris M, Cruz-Romero M, Cummins E (2014). Evaluation and simulation of silver and copper nanoparticle migration from polyethylene nanocomposites to food and an associated exposure assessment. Journal of Agricultural and Food Chemistry 62: 1403–1411. DOI 10.1021/jf404038y. [Google Scholar] [CrossRef]
Dawood MA, Eweedah NM, Moustafa EM, El-Sharawy ME, Soliman AA, Amer AA, Atia MH (2020). Copper nanoparticles mitigate the growth, immunity, and oxidation resistance in common carp (Cyprinus carpio). Biological Trace Element Research 198: 1–10. DOI 10.1007/s12011-020-02068-0. [Google Scholar] [CrossRef]
de Bie P, Muller P, Wijmenga C, Klomp LW (2007). Molecular pathogenesis of Wilson and Menkes disease: Correlation of mutations with molecular defects and disease phenotypes. Journal of Medical Genetics 44: 673–688. DOI 10.1136/jmg.2007.052746. [Google Scholar] [CrossRef]
Dekkers S, Bouwmeester H, Bos PM, Peters RJ, Rietveld AG, Oomen AG (2013). Knowledge gaps in risk assessment of nanosilica in food: Evaluation of the dissolution and toxicity of different forms of silica. Nanotoxicology 7: 367–377. DOI 10.3109/17435390.2012.662250. [Google Scholar] [CrossRef]
Du J, Fu L, Li H, Xu S, Zhou Q, Tang J (2019). The potential hazards and ecotoxicity of CuO nanoparticles: An overview. Toxin Reviews 40: 1–13. DOI 10.1080/15569543.2019.1670211. [Google Scholar] [CrossRef]
Duan H, Yu L, Tian F, Zhai Q, Fan L, Chen W (2020). Gut microbiota: A target for heavy metal toxicity and a probiotic protective strategy. Science of the Total Environment 742: 140429. DOI 10.1016/j.scitotenv.2020.140429. [Google Scholar] [CrossRef]
El-Kassas S, Abdo SE, El-Naggar K, Abdo W, Kirrella AA, Nashar TO (2018). Ameliorative effect of dietary supplementation of copper oxide nanoparticles on inflammatory and immune reponses in commercial broiler under normal and heat-stress housing conditions. Journal of Thermal Biology 78: 235–246. DOI 10.1016/j.jtherbio.2018.10.009. [Google Scholar] [CrossRef]
El-Katcha MI, Soltan MA, Khalifa E, Fadl SE, Hassan A, El-Shimey OK (2020). Growth and immune response of broiler chicks fed on oxidized oil containing diets and supplemented with different copper sources and levels. Alexandria Journal for Veterinary Sciences 66: 15–26. DOI 10.5455/ajvs.98495. [Google Scholar] [CrossRef]
Elkloub K, El Moustafa M, Ghazalah A, Rehan A (2015). Effect of dietary nanosilver on broiler performance. International Journal of Poultry Science 14: 177–182. DOI 10.3923/ijps.2015.177.182. [Google Scholar] [CrossRef]
Eng SK, Pusparajah P, Ab Mutalib NS, Ser HL, Chan KG, Lee LH (2015). Salmonella: A review on pathogenesis, epidemiology and antibiotic resistance. Frontiers in Life Science 8: 284–293. DOI 10.1080/21553769.2015.1051243. [Google Scholar] [CrossRef]
Fàbrega A, Vila J (2013). Salmonella enterica serovar Typhimurium skills to succeed in the host: Virulence and regulation. Clinical Microbiology Reviews 26: 308–341. DOI 10.1128/CMR.00066-12. [Google Scholar] [CrossRef]
Faizan M, Faraz A, Hayat S, Bhat JA, Yu F (2021). Zinc oxide nanoparticles and epibrassinolide enhanced growth of tomato via modulating antioxidant activity and photosynthetic performance. BIOCELL 45: 1081–1093. DOI 10.32604/biocell.2021.015363. [Google Scholar] [CrossRef]
Falony G, Vieira-Silva S, Raes J (2018). Richness and ecosystem development across faecal snapshots of the gut microbiota. Nature Microbiology 3: 526–528. DOI 10.1038/s41564-018-0143-5. [Google Scholar] [CrossRef]
Fan Y, Pedersen O (2020). Gut microbiota in human metabolic health and disease. Nature Reviews Microbiology 19: 1–17. DOI 10.1038/s41579-020-0433-9. [Google Scholar] [CrossRef]
Fazeli M, Hassanzadeh P, Alaei S (2011). Cadmium chloride exhibits a profound toxic effect on bacterial microflora of the mice gastrointestinal tract. Human Experimental Toxicology 30: 152–159. DOI 10.1177/0960327110369821. [Google Scholar] [CrossRef]
Festa RA, Thiele DJ (2011). Copper: An essential metal in biology. Current Biology 21: R877–R883. DOI 10.1016/j.cub.2011.09.040. [Google Scholar] [CrossRef]
Fondevila M, Herrer R, Casallas MC, Abecia L, Ducha JJ (2009). Silver nanoparticles as a potential antimicrobial additive for weaned pigs. Animal Feed Science and Technology 150: 259–269. DOI 10.1016/j.anifeedsci.2008.09.003. [Google Scholar] [CrossRef]
Foster JA, Neufeld KAM (2013). Gut-brain axis: How the microbiome influences anxiety and depression. Trends in Neurosciences 36: 305–312. DOI 10.1016/j.tins.2013.01.005. [Google Scholar] [CrossRef]
Fröhlich EE, Fröhlich E (2016). Cytotoxicity of nanoparticles contained in food on intestinal cells and the gut microbiota. International Journal of Molecular Sciences 17: 509. DOI 10.3390/ijms17040509. [Google Scholar] [CrossRef]
Fujisaka S, Ussar S, Clish C, Devkota S, Dreyfuss JM, Sakaguchi M, Soto M, Konishi M, Softic S, Altindis E (2016). Antibiotic effects on gut microbiota and metabolism are host dependent. The Journal of Clinical Investigation 126: 4430–4443. DOI 10.1172/JCI86674. [Google Scholar] [CrossRef]
Gangadoo S, Dinev I, Chapman J, Hughes RJ, Van TTH, Moore RJ, Stanley D (2018). Selenium nanoparticles in poultry feed modify gut microbiota and increase abundance of Faecalibacterium prausnitzii. Applied Microbiology and Biotechnology 102: 1455–1466. DOI 10.1007/s00253-017-8688-4. [Google Scholar] [CrossRef]
Gangadoo S, Stanley D, Hughes RJ, Moore RJ, Chapman J (2016). Nanoparticles in feed: Progress and prospects in poultry research. Trends in Food Science & Technology 58: 115–126. DOI 10.1016/j.tifs.2016.10.013. [Google Scholar] [CrossRef]
García-Ruiz A, Crespo J, López-de-Luzuriaga J, Olmos M, Monge M, Rodríguez-Álfaro M, Martín-Álvarez P, Bartolome B, Moreno-Arribas M (2015). Novel biocompatible silver nanoparticles for controlling the growth of lactic acid bacteria and acetic acid bacteria in wines. Food Control 50: 613–619. DOI 10.1016/j.foodcont.2014.09.035. [Google Scholar] [CrossRef]
Gevers D, Kugathasan S, Denson LA, Vázquez-Baeza Y, Van Treuren W, Ren B, Schwager E, Knights D, Song SJ, Yassour M (2014). The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host & Microbe 15: 382–392. DOI 10.1016/j.chom.2014.02.005. [Google Scholar] [CrossRef]
Ghebretatios M, Schaly S, Prakash S (2021). Nanoparticles in the food industry and their impact on human gut microbiome and diseases. International Journal of Molecular Sciences 22: 1942. DOI 10.3390/ijms22041942. [Google Scholar] [CrossRef]
Gil-Sánchez I, Cueva C, Sanz-Buenhombre M, Guadarrama A, Moreno-Arribas MV, Bartolomé B (2018). Dynamic gastrointestinal digestion of grape pomace extracts: Bioaccessible phenolic metabolites and impact on human gut microbiota. Journal of Food Composition and Analysis 68: 41–52. DOI 10.1016/j.jfca.2017.05.005. [Google Scholar] [CrossRef]
Gong J, Si W, Forster RJ, Huang R, Yu H, Yin Y, Yang C, Han Y (2007). 16S rRNA gene-based analysis of mucosa-associated bacterial community and phylogeny in the chicken gastrointestinal tracts: From crops to ceca. FEMS Microbiology Ecology 59: 147–157. DOI 10.1111/j.1574-6941.2006.00193.x. [Google Scholar] [CrossRef]
Gonzalez P, Baudrimont M, Boudou A, Bourdineaud JP (2006). Comparative effects of direct cadmium contamination on gene expression in gills, liver, skeletal muscles and brain of the zebrafish (Danio rerio). Biometals 19: 225–235. DOI 10.1007/s10534-005-5670-x. [Google Scholar] [CrossRef]
Gophna U, Konikoff T, Nielsen HB (2017). Oscillospira and related bacteria–From metagenomic species to metabolic features. Environmental Microbiology 19: 835–841. DOI 10.1111/1462-2920.13658. [Google Scholar] [CrossRef]
Gupta A, Landis RF, Rotello VM (2016). Nanoparticle-based antimicrobials: Surface functionality is critical. F1000Research 5: 364. DOI 10.12688/f1000research.7595.1. [Google Scholar] [CrossRef]
Gupta A, Lutsenko S (2009). Human copper transporters: Mechanism, role in human diseases and therapeutic potential. Future Medicinal Chemistry 1: 1125–1142. DOI 10.4155/fmc.09.84. [Google Scholar] [CrossRef]
Gupta A, Saleh NM, Das R, Landis RF, Bigdeli A, Motamedchaboki K, Campos AR, Pomeroy K, Mahmoudi M, Rotello VM (2017). Synergistic antimicrobial therapy using nanoparticles and antibiotics for the treatment of multidrug-resistant bacterial infection. Nano Futures 1: 015004. DOI 10.1088/2399-1984/aa69fb. [Google Scholar] [CrossRef]
Guyard-Nicodeme M, Keita A, Quesne S, Amelot M, Poezevara T, Le Berre B, Sánchez J, Vesseur P, Martín Á, Medel P (2016). Efficacy of feed additives against Campylobacter in live broilers during the entire rearing period. Poultry Science 95: 298–305. DOI 10.3382/ps/pev303. [Google Scholar] [CrossRef]
Handy RD, Owen R, Valsami-Jones E (2008). The ecotoxicology of nanoparticles and nanomaterials: current status, knowledge gaps, challenges, and future needs. Ecotoxicology 17: 315–325. DOI 10.1007/s10646-008-0206-0. [Google Scholar] [CrossRef]
Hansson GK, Jonasson L (2009). The discovery of cellular immunity in the atherosclerotic plaque. Arteriosclerosis, Thrombosis, Vascular Biology 29: 1714–1717. DOI 10.1161/ATVBAHA.108.179713. [Google Scholar] [CrossRef]
Heiss CN, Olofsson LE (2019). The role of the gut microbiota in development, function and disorders of the central nervous system and the enteric nervous system. Journal of Neuroendocrinology 3: e12684. DOI 10.1111/jne.12684. [Google Scholar] [CrossRef]
Hill DA, Siracusa MC, Abt MC, Kim BS, Kobuley D, Kubo M, Kambayashi T, LaRosa DF, Renner ED, Orange JS (2012). Commensal bacteria-derived signals regulate basophil hematopoiesis and allergic inflammation. Nature Medicine 18: 538–546. DOI 10.1038/nm.2657. [Google Scholar] [CrossRef]
Ho KT, Portis L, Chariton AA, Pelletier M, Cantwell M, Katz D, Cashman M, Parks A, Baguley JG, Conrad-Forrest N (2018). Effects of micronized and nano-copper azole on marine benthic communities. Environmental Toxicology and Chemistry 37: 362–375. DOI 10.1002/etc.3954. [Google Scholar] [CrossRef]
Huo S, Jiang Y, Gupta A, Jiang Z, Landis RF, Hou S, Liang XJ, Rotello VM (2016). Fully zwitterionic nanoparticle antimicrobial agents through tuning of core size and ligand structure. ACS Nano 10: 8732–8737. DOI 10.1021/acsnano.6b04207. [Google Scholar] [CrossRef]
Hurdle JG, O’neill AJ, Chopra I, Lee RE (2011). Targeting bacterial membrane function: An underexploited mechanism for treating persistent infections. Nature Reviews Microbiology 9: 62–75. DOI 10.1038/nrmicro2474. [Google Scholar] [CrossRef]
Huttenhower C, Gevers D, Knight R, Abubucker S, Badger JH, Chinwalla AT, Creasy HH, Earl AM, FitzGerald MG, Fulton RS (2012). Structure, function and diversity of the healthy human microbiome. Nature 486: 207–214. DOI 10.1038/nature11234. [Google Scholar] [CrossRef]
Hyland NP, Cryan JF (2016). Microbe-host interactions: Influence of the gut microbiota on the enteric nervous system. Developmental Biology 417: 182–187. DOI 10.1016/j.ydbio.2016.06.027. [Google Scholar] [CrossRef]
Jandhyala SM, Talukdar R, Subramanyam C, Vuyyuru H, Sasikala M, Reddy DN (2015). Role of the normal gut microbiota. World Journal of Gastroenterology 21: 8787. DOI 10.3748/wjg.v21.i29.8787. [Google Scholar] [CrossRef]
Jarosz Ł., Marek A, Grądzki Z, Kwiecień M, Kaczmarek B (2018). The effect of feed supplementation with a copper-glycine chelate and copper sulphate on selected humoral and cell-mediated immune parameters, plasma superoxide dismutase activity, ceruloplasmin and cytokine concentration in broiler chickens. Journal of Animal Physiology and Animal Nutrition 102: e326–e336. DOI 10.1111/jpn.12750. [Google Scholar] [CrossRef]
Jie Z, Xia H, Zhong SL, Feng Q, Li S, Liang S, Zhong H, Liu Z, Gao Y, Zhao H (2017). The gut microbiome in atherosclerotic cardiovascular disease. Nature Communications 8: 1–12. DOI 10.1038/s41467-017-00900-1. [Google Scholar] [CrossRef]
Jin M, Qian Z, Yin J, Xu W, Zhou X (2019). The role of intestinal microbiota in cardiovascular disease. Journal of Cellular and Molecular Medicine 23: 2343–2350. DOI 10.1111/jcmm.14195. [Google Scholar] [CrossRef]
Kamada N, Seo SU, Chen GY, Núñez G (2013). Role of the gut microbiota in immunity and inflammatory disease. Nature Reviews Immunology 13: 321–335. DOI 10.1038/nri3430. [Google Scholar] [CrossRef]
Kamaruzzaman NF, Kendall S, Good L (2017). Targeting the hard to reach: Challenges and novel strategies in the treatment of intracellular bacterial infections. British Journal of Pharmacology 174: 2225–2236. DOI 10.1111/bph.13664. [Google Scholar] [CrossRef]
Karavolos M, Holban A (2016). Nanosized drug delivery systems in gastrointestinal targeting: Interactions with microbiota. Pharmaceuticals 9: 62. DOI 10.3390/ph9040062. [Google Scholar] [CrossRef]
Karlsson FH, Fåk F, Nookaew I, Tremaroli V, Fagerberg B, Petranovic D, Bäckhed F, Nielsen J (2012). Symptomatic atherosclerosis is associated with an altered gut metagenome. Nature Communications 3: 1–8. DOI 10.1038/ncomms2266. [Google Scholar] [CrossRef]
Kawase T, Nagasawa M, Ikeda H, Yasuo S, Koga Y, Furuse M (2017). Gut microbiota of mice putatively modifies amino acid metabolism in the host brain. British Journal of Nutrition 117: 775–783. DOI 10.1017/S0007114517000678. [Google Scholar] [CrossRef]
Khan BA, Karim F, Khan MK, Haider F, Khan S (2021). Synthesis and characterization of polymeric responsive CMC/Pectin hydrogel films loaded with Tamarix aphylla extract as potential wound dressings. BIOCELL 45: 1273–1285. DOI 10.32604/biocell.2021.015323. [Google Scholar] [CrossRef]
Kim E, Lembert MM, Opiyo SO, Ahmer BM, Cormet-Boyaka E, Boyaka PN (2017). Differential role of oxidative stress pathways and microbiota in the development of allergen specific IgE following chronic ingestion of low doses of cadmium. The Journal of Immunology 198. [Google Scholar]
Kim E, Xu X, Steiner H, Ahmer B, Cormet-Boyaka E, Boyaka P (2015). Chronic ingestion of low doses of cadmium alters the gut microbiome and immune homeostasis to enhance allergic sensitization (MUC9P.743). The Journal of Immunology 194. [Google Scholar]
Korem T, Zeevi D, Suez J, Weinberger A, Avnit-Sagi T, Pompan-Lotan M, Matot E, Jona G, Harmelin A, Cohen N (2015). Growth dynamics of gut microbiota in health and disease inferred from single metagenomic samples. Science 349: 1101–1106. DOI 10.1126/science.aac4812. [Google Scholar] [CrossRef]
Kumar S, Sharma A (2019). Cadmium toxicity: Effects on human reproduction and fertility. Reviews on Environmental Health 34: 327–338. DOI 10.1515/reveh-2019-0016. [Google Scholar] [CrossRef]
Lai X, Wei Y, Zhao H, Chen S, Bu X, Lu F, Qu D, Yao L, Zheng J, Zhang J (2015). The effect of Fe2O3 and ZnO nanoparticles on cytotoxicity and glucose metabolism in lung epithelial cells. Journal of Applied Toxicology 35: 651–664. DOI 10.1002/jat.3128. [Google Scholar] [CrossRef]
Lamas B, Martins Breyner N, Houdeau E (2020). Impacts of foodborne inorganic nanoparticles on the gut microbiota-immune axis: Potential consequences for host health. Particle Fibre Toxicology 17: 1–22. DOI 10.1186/s12989-020-00349-z. [Google Scholar] [CrossRef]
Lanter BB, Sauer K, Davies DG (2014). Bacteria present in carotid arterial plaques are found as biofilm deposits which may contribute to enhanced risk of plaque rupture. mBio 5: 3149. DOI 10.1128/mBio.01206-14. [Google Scholar] [CrossRef]
Le Chatelier E, Nielsen T, Qin J, Prifti E, Hildebrand F, Falony G, Almeida M, Arumugam M, Batto JM, Kennedy S (2013). Richness of human gut microbiome correlates with metabolic markers. Nature 500: 541–546. DOI 10.1038/nature12506. [Google Scholar] [CrossRef]
Ley RE, Turnbaugh PJ, Klein S, Gordon J (2006). Human gut microbes associated with obesity. Nature Reviews Microbiology 444: 1022–1023. DOI 10.1038/4441022a. [Google Scholar] [CrossRef]
Li J, Tang M, Xue Y (2019). Review of the effects of silver nanoparticle exposure on gut bacteria. Journal of Applied Toxicology 39: 27–37. DOI 10.1002/jat.3729. [Google Scholar] [CrossRef]
Lin Y, Huang J, Li M, Cheng C, Lien T (2015). Effects of supplemental nanoparticle trivalent chromium on the nutrient utilization, growth performance and serum traits of broilers. Journal of Animal Physiology and Animal Nutrition 99: 59–65. DOI 10.1111/jpn.12215. [Google Scholar] [CrossRef]
Liu R, Hong J, Xu X, Feng Q, Zhang D, Gu Y, Shi J, Zhao S, Liu W, Wang X (2017). Gut microbiome and serum metabolome alterations in obesity and after weight-loss intervention. Nature Medicine 23: 859–868. DOI 10.1038/nm.4358. [Google Scholar] [CrossRef]
Liu Y, Li Y, Xia Y, Liu K, Ren L, Ji Y (2020). The dysbiosis of gut microbiota caused by low-dose cadmium aggravate the injury of mice liver through increasing intestinal permeability. Microorganisms 8: 211. DOI 10.3390/microorganisms8020211. [Google Scholar] [CrossRef]
Lorincz MT (2018). Wilson disease and related copper disorders, Handbook of Clinical Neurology, vol. 147, pp. 279–292. Elsevier. [Google Scholar]
Lutsenko S, Barnes NL, Bartee MY, Dmitriev OY (2007). Function and regulation of human copper-transporting ATPases. Physiological Reviews 87: 1011–1046. DOI 10.1152/physrev.00004.2006. [Google Scholar] [CrossRef]
Lynch SV, Pedersen O (2016). The human intestinal microbiome in health and disease. New England Journal of Medicine 375: 2369–2379. DOI 10.1056/NEJMra1600266. [Google Scholar] [CrossRef]
Makabenta JMV, Nabawy A, Li CH, Schmidt-Malan S, Patel R, Rotello VM (2021). Nanomaterial-based therapeutics for antibiotic-resistant bacterial infections. Nature Reviews Microbiology 19: 1–14. DOI 10.1038/s41579-020-0420-1. [Google Scholar] [CrossRef]
Matsuoka K, Kanai T (2015). The gut microbiota and inflammatory bowel disease. In: Seminars in Immunopathology, vol. 37, no. 1, pp. 47–55. Springer Berlin Heidelberg. [Google Scholar]
McAllister EJ, Dhurandhar NV, Keith SW, Aronne LJ, Barger J, Baskin M, Benca RM, Biggio J, Boggiano MM, Eisenmann JC (2009). Ten putative contributors to the obesity epidemic. Critical Reviews in Food Science and Nutrition 49: 868–913. DOI 10.1080/10408390903372599. [Google Scholar] [CrossRef]
McClements DJ, Xiao H (2017). Is nano safe in foods? Establishing the factors impacting the gastrointestinal fate and toxicity of organic and inorganic food-grade nanoparticles. npj Science of Food 1: 1–13. DOI 10.1038/s41538-017-0005-1. [Google Scholar] [CrossRef]
McVey Neufeld K, Mao Y, Bienenstock J, Foster J, Kunze W (2013). The microbiome is essential for normal gut intrinsic primary afferent neuron excitability in the mouse. Neurogastroenterology and Motility 25: 183–e188. DOI 10.1111/nmo.12049. [Google Scholar] [CrossRef]
Memar MY, Ghotaslou R, Samiei M, Adibkia K (2018). Antimicrobial use of reactive oxygen therapy: Current insights. Infection and Drug Resistance 11: 567–576. DOI 10.2147/IDRS142397. [Google Scholar] [CrossRef]
Miller KP, Wang L, Benicewicz BC, Decho AW (2015). Inorganic nanoparticles engineered to attack bacteria. Chemical Society Reviews 44: 7787–7807. DOI 10.1039/C5CS00041F. [Google Scholar] [CrossRef]
Miller TL, Wolin M, de Macario EC, Macario A (1982). Isolation of Methanobrevibacter smithii from human feces. Applied and Environmental Microbiology 43: 227–232. DOI 10.1128/aem.43.1.227-232.1982. [Google Scholar] [CrossRef]
Mocchegiani E, Santarelli L, Muzzioli M, Fabris N (1995). Reversibility of the thymic involution and of age-related peripheral immune dysfunctions by zinc supplementation in old mice. International Journal of Immunopharmacology 17: 703–718. DOI 10.1016/0192-0561(95)00059-B. [Google Scholar] [CrossRef]
Monge M, Moreno-Arribas MV (2016). Applications of nanotechnology in wine production and quality and safety control. In: Wine Safety, Consumer Preference, and Human Health, pp. 51-69. Springer. [Google Scholar]
Naseribafrouei A, Hestad K, Avershina E, Sekelja M, Linløkken A, Wilson R, Rudi K (2014). Correlation between the human fecal microbiota and depression. Neurogastroenterology Motility 26: 1155–1162. DOI 10.1111/nmo.12378. [Google Scholar] [CrossRef]
Novakovic M, Rout A, Kingsley T, Kirchoff R, Singh A, Verma V, Kant R, Chaudhary R (2020). Role of gut microbiota in cardiovascular diseases. World Journal of Cardiology 12: 110–122. DOI 10.4330/wjc.v12.i4.110. [Google Scholar] [CrossRef]
Obeng AS, Rickard H, Ndi O, Sexton M, Barton M (2012). Antibiotic resistance, phylogenetic grouping and virulence potential of Escherichia coli isolated from the faeces of intensively farmed and free range poultry. Veterinary Microbiology 154: 305–315. DOI 10.1016/j.vetmic.2011.07.010. [Google Scholar] [CrossRef]
Ofosu A (2016). Clostridium difficile infection: A review of current and emerging therapies. Annals of Gastroenterology: Quarterly Publication of the Hellenic Society of Gastroenterology 29: 147–154. DOI 10.20524/aog.2016.0006. [Google Scholar] [CrossRef]
Ognik K, Sembratowicz I, Cholewińska E, Wlazło Ł, Nowakowicz-Dębek B, Szlązak R, Tutaj K (2016). The effect of chemically-synthesized silver nanoparticles on performance and the histology and microbiological Profile of the jejunum in chickens. Annals of Animal Science 16: 439–450. DOI 10.1515/aoas-2015-0067. [Google Scholar] [CrossRef]
World Health Organization (2000). Obesity: Preventing and Managing the Global Epidemic: Report on a WHO Consultation (WHO Technical Report Series 894). Geneva, Switzerland: World Health Organization. [Google Scholar]
World Health Organization (2017). Obesity and overweight. http://www.who.int/mediacentre/factsheets/fs311/en/. [Google Scholar]
Patel P, Kansara K, Senapati VA, Shanker R, Dhawan A, Kumar A (2016). Cell cycle dependent cellular uptake of zinc oxide nanoparticles in human epidermal cells. Mutagenesis 31: 481–490. DOI 10.1093/mutage/gew014. [Google Scholar] [CrossRef]
Tyler Patterson T, Grandhi R (2020). Gut microbiota and neurologic diseases and injuries. In: Chen, P. (ed.Gut Microbiota and Pathogenesis of Organ Injury. Advances in Experimental Medicine and Biology, vol. 1238. Springer, Singapore. DOI 10.1007/978-981-15-2385-4_6. [Google Scholar] [CrossRef]
Pelgrift RY, Friedman AJ (2013). Nanotechnology as a therapeutic tool to combat microbial resistance. Advanced Drug Delivery Reviews 65: 1803–1815. DOI 10.1016/j.addr.2013.07.011. [Google Scholar] [CrossRef]
Pietroiusti A, Magrini A, Campagnolo L (2016). New frontiers in nanotoxicology: Gut microbiota/microbiome-mediated effects of engineered nanomaterials. Toxicology and Applied Pharmacology 299: 90–95. DOI 10.1016/j.taap.2015.12.017. [Google Scholar] [CrossRef]
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T (2010). A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464: 59–65. DOI 10.1038/nature08821. [Google Scholar] [CrossRef]
Qiu K, Durham PG, Anselmo AC (2018). Inorganic nanoparticles and the microbiome. Nano Research 11: 4936–4954. DOI 10.1007/s12274-018-2137-2. [Google Scholar] [CrossRef]
Reed S, Neuman H, Moscovich S, Glahn RP, Koren O, Tako E (2015). Chronic zinc deficiency alters chick gut microbiota composition and function. Nutrients 7: 9768–9784. DOI 10.3390/nu7125497. [Google Scholar] [CrossRef]
Rosenfeld ME, Campbell LA (2011). Pathogens and atherosclerosis: Update on the potential contribution of multiple infectious organisms to the pathogenesis of atherosclerosis. Thrombosis Haemostasis 106: 858–867. DOI 10.1160/TH11-06-0392. [Google Scholar] [CrossRef]
Ross AC, Caballero B, Cousins RJ, Tucker KL. (2020). Modern Nutrition in Health and Disease. Jones & Bartlett Learning: Lippincott Williams & Wilkins. [Google Scholar]
Rothschild D, Weissbrod O, Barkan E, Kurilshikov A, Korem T, Zeevi D, Costea PI, Godneva A, Kalka IN, Bar N (2018). Environment dominates over host genetics in shaping human gut microbiota. Nature 555: 210–215. DOI 10.1038/nature25973. [Google Scholar] [CrossRef]
Saptarshi SR, Duschl A, Lopata AL (2015). Biological reactivity of zinc oxide nanoparticles with mammalian test systems: An overview. Nanomedicine: Nanotechnology, Biology, and Medicine 10: 2075–2092. DOI 10.2217/nnm.15.44. [Google Scholar] [CrossRef]
Sawosz E, Binek M, Grodzik M, Zielińska M, Sysa P, Szmidt M, Niemiec T, Chwalibog A (2007). Influence of hydrocolloidal silver nanoparticles on gastrointestinal microflora and morphology of enterocytes of quails. Archives of Animal Nutrition 61: 444–451. DOI 10.1080/17450390701664314. [Google Scholar] [CrossRef]
Scanlan PD, Shanahan F, O’Mahony C, Marchesi JR (2006). Culture-independent analyses of temporal variation of the dominant fecal microbiota and targeted bacterial subgroups in Crohn’s disease. Journal of Clinical Microbiology 44: 3980–3988. DOI 10.1128/JCM.00312-06. [Google Scholar] [CrossRef]
Schwabe RF, Jobin C (2013). The microbiome and cancer. Nature Reviews Cancer 13: 800–812. DOI 10.1038/nrc3610. [Google Scholar] [CrossRef]
Scott A, Vadalasetty K, Łukasiewicz M, Jaworski S, Wierzbicki M, Chwalibog A, Sawosz E (2018a). Effect of different levels of copper nanoparticles and copper sulphate on performance, metabolism and blood biochemical profiles in broiler chicken. Journal of Animal Physiology and Animal Nutrition 102: e364–e373. DOI 10.1111/jpn.12754. [Google Scholar] [CrossRef]
Scott A, Vadalasetty KP, Chwalibog A, Sawosz E (2018b). Copper nanoparticles as an alternative feed additive in poultry diet: A review. Nanotechnology Reviews 7: 69–93. DOI 10.1515/ntrev-2017-0159. [Google Scholar] [CrossRef]
Sergeevna AS, Anatolevich SO, Nurchallaeva SG, Aleksandrovich BA, Petrovich RE, Vladimirovich SS, Alexeevna GM, Diadorovitch II (2018). Silver in the meat and organs of broiler chickens in case of using colloidal silver as an alternative to antibiotics. BioMetals 31: 975–980. DOI 10.1007/s10534-018-0141-3. [Google Scholar] [CrossRef]
Shamaila S, Zafar N, Riaz S, Sharif R, Nazir J, Naseem S (2016). Gold nanoparticles: An efficient antimicrobial agent against enteric bacterial human pathogen. Nanomaterials 6: 71. DOI 10.3390/nano6040071. [Google Scholar] [CrossRef]
Shao Y, Lei Z, Yuan J, Yang Y, Guo Y, Zhang B (2014). Effect of zinc on growth performance, gut morphometry, and cecal microbial community in broilers challenged with Salmonella enterica serovar typhimurium. Journal of Microbiology 52: 1002–1011. DOI 10.1007/s12275-014-4347-y. [Google Scholar] [CrossRef]
Silvan JM, Zorraquin-Peña I, Gonzalez de Llano D, Moreno-Arribas M, Martinez-Rodriguez AJ (2018). Antibacterial activity of glutathione-stabilized silver nanoparticles against Campylobacter multidrug-resistant strains. Frontiers in Microbiology 9: 458. DOI 10.3389/fmicb.2018.00458. [Google Scholar] [CrossRef]
Sirelkhatim A, Mahmud S, Seeni A, Kaus NHM, Ann LC, Bakhori SKM, Hasan H, Mohamad D (2015). Review on zinc oxide nanoparticles: Antibacterial activity and toxicity mechanism. Nano-Micro Letters 7: 219–242. DOI 10.1007/s40820-015-0040-x. [Google Scholar] [CrossRef]
Soenen SJ, Rivera-Gil P, Montenegro JM, Parak WJ, de Smedt SC, Braeckmans K (2011). Cellular toxicity of inorganic nanoparticles: Common aspects and guidelines for improved nanotoxicity evaluation. Nano Today 6: 446–465. DOI 10.1016/j.nantod.2011.08.001. [Google Scholar] [CrossRef]
Sommer F, Bäckhed F (2013). The gut microbiota—Masters of host development and physiology. Nature Reviews Microbiology 11: 227–238. DOI 10.1038/nrmicro2974. [Google Scholar] [CrossRef]
Song W, Anselmo AC, Huang L (2019). Nanotechnology intervention of the microbiome for cancer therapy. Nature Nanotechnology 14: 1093–1103. DOI 10.1038/s41565-019-0589-5. [Google Scholar] [CrossRef]
Stecher B, Hardt WD (2008). The role of microbiota in infectious disease. Trends in Microbiology 16: 107–114. DOI 10.1016/j.tim.2007.12.008. [Google Scholar] [CrossRef]
Sun N, Wang H, Ju Z, Zhao H (2018). Effects of chronic cadmium exposure on metamorphosis, skeletal development, and thyroid endocrine disruption in Chinese toad Bufo gargarizans tadpoles. Environmental Toxicology Chemistry 37: 213–223. DOI 10.1002/etc.3947. [Google Scholar] [CrossRef]
Surai P (2002a). Selenium in poultry nutrition 1. Antioxidant properties, deficiency and toxicity. World’s Poultry Science Journal 58: 333–347. DOI 10.1079/WPS20020026. [Google Scholar] [CrossRef]
Surai P (2002b). Selenium in poultry nutrition 2. Reproduction, egg and meat quality and practical applications. World’s Poultry Science Journal 58: 431–450. DOI 10.1079/WPS20020032. [Google Scholar] [CrossRef]
Suzuki K, Ogra Y (2002). Metabolic pathway for selenium in the body: Speciation by HPLC-ICP MS with enriched Se. Food Additives & Contaminants 19: 974–983. DOI 10.1080/02652030210153578. [Google Scholar] [CrossRef]
Swart E, Dvorak J, Hernádi S, Goodall T, Kille P, Spurgeon D, Svendsen C, Prochazkova P (2020). The effects of in vivo exposure to copper oxide nanoparticles on the gut microbiome, host immunity, and susceptibility to a bacterial infection in earthworms. Nanomaterials 10: 1337. DOI 10.3390/nano10071337. [Google Scholar] [CrossRef]
Takaishi H, Matsuki T, Nakazawa A, Takada T, Kado S, Asahara T, Kamada N, Sakuraba A, Yajima T, Higuchi H (2008). Imbalance in intestinal microflora constitution could be involved in the pathogenesis of inflammatory bowel disease. International Journal of Medical Microbiology 298: 463–472. DOI 10.1016/j.ijmm.2007.07.016. [Google Scholar] [CrossRef]
Thingholm LB, Rühlemann MC, Koch M, Fuqua B, Laucke G, Boehm R, Bang C, Franzosa EA, Hübenthal M, Rahnavard A (2019). Obese individuals with and without type 2 diabetes show different gut microbial functional capacity and composition. Cell Host & Microbe 26: 252-264. e210. DOI 10.1016/j.chom.2019.07.004. [Google Scholar] [CrossRef]
Timbo BB, Ross MP, McCarthy PV, Lin CTJ (2006). Dietary supplements in a national survey: Prevalence of use and reports of adverse events. Journal of the American Dietetic Association 106: 1966–1974. DOI 10.1016/j.jada.2006.09.002. [Google Scholar] [CrossRef]
Tims S, Derom C, Jonkers DM, Vlietinck R, Saris WH, Kleerebezem M, de Vos WM, Zoetendal EG (2013). Microbiota conservation and BMI signatures in adult monozygotic twins. The ISME Journal 7: 707–717. DOI 10.1038/ismej.2012.146. [Google Scholar] [CrossRef]
Tong M, Li X, Parfrey LW, Roth B, Ippoliti A, Wei B, Borneman J, McGovern DP, Frank DN, Li E (2013). A modular organization of the human intestinal mucosal microbiota and its association with inflammatory bowel disease. PLoS One 8: e80702. DOI 10.1371/journal.pone.0080702. [Google Scholar] [CrossRef]
Torok VA, Allison GE, Percy NJ, Ophel-Keller K, Hughes RJ (2011). Influence of antimicrobial feed additives on broiler commensal posthatch gut microbiota development and performance. Applied and Environmental Microbiology 77: 3380–3390. DOI 10.1128/AEM.02300-10. [Google Scholar] [CrossRef]
Tremlett H, Bauer KC, Appel-Cresswell S, Finlay BB, Waubant E (2017). The gut microbiome in human neurological disease: A review. Annals of Neurology 81: 369–382. DOI 10.1002/ana.24901. [Google Scholar] [CrossRef]
Tümer Z, Møller LB (2010). Menkes disease. European Journal of Human Genetics 18: 511–518. DOI 10.1038/ejhg.2009.187. [Google Scholar] [CrossRef]
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI (2006). An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444: 1027–1031. DOI 10.1038/nature05414. [Google Scholar] [CrossRef]
Tyakht AV, Kostryukova ES, Popenko AS, Belenikin MS, Pavlenko AV, Larin AK, Karpova IY, Selezneva OV, Semashko TA, Ospanova EA (2013). Human gut microbiota community structures in urban and rural populations in Russia. Nature Communications 4: 1–9. DOI 10.1038/ncomms3469. [Google Scholar] [CrossRef]
Ursell LK, Metcalf JL, Parfrey LW, Knight R (2012). Defining the human microbiome. Nutrition Reviews 70: S38–S44. DOI 10.1111/j.1753-4887.2012.00493.x. [Google Scholar] [CrossRef]
Utembe W, Tlotleng N, Kamng’ona A (2022). A systematic review on the effects of nanomaterials on gut microbiota. Current Research in Microbial Sciences 3: 100118. DOI 10.1016/j.crmicr.2022.100118. [Google Scholar] [CrossRef]
Vickers NJ (2017). Animal communication: When i’m calling you, will you answer too? Current Biology 27: R713–R715. DOI 10.1016/j.cub.2017.05.064. [Google Scholar] [CrossRef]
Vieira de Souza F, Roque R, Silva Moreira J, Resende de Souza M, Nicoli J, Neumann E, Cantini Nunes Á (2012). Transfer of antibiotic resistance determinants between lactobacilli isolates from the gastrointestinal tract of chicken. Beneficial Microbes 3: 137–144. DOI 10.3920/BM2011.0058. [Google Scholar] [CrossRef]
Walczak AP, Fokkink R, Peters R, Tromp P, Herrera Rivera ZE, Rietjens IM, Hendriksen PJ, Bouwmeester H (2012). Behaviour of silver nanoparticles and silver ions in an in vitro human gastrointestinal digestion model. Nanotoxicology 7: 1198–1210. DOI 10.3109/17435390.2012.726382. [Google Scholar] [CrossRef]
Wang C, Wang M, Ye S, Tao W, Du Y (2011a). Effects of copper-loaded chitosan nanoparticles on growth and immunity in broilers. Poultry Science 90: 2223–2228. DOI 10.3382/ps.2011-01511. [Google Scholar] [CrossRef]
Wang L, Xu T, Lei WW, Liu DM, Li YJ, Xuan RJ, Ma JJ (2011b). Cadmium-induced oxidative stress and apoptotic changes in the testis of freshwater crab, Sinopotamon henanense. PLoS One 6: e27853. DOI 10.1371/journal.pone.0027853. [Google Scholar] [CrossRef]
Wang N, Tan HY, Li S, Xu Y, Guo W, Feng Y (2017). Supplementation of micronutrient selenium in metabolic diseases: Its role as an antioxidant. Oxidative Medicine and Cellular Longevity 2017: 1–13. DOI 10.1155/2017/7478523. [Google Scholar] [CrossRef]
Wang W, Chen L, Zhou R, Wang X, Song L, Huang S, Wang G, Xia B (2014). Increased proportions of bifidobacterium and the lactobacillus group and loss of butyrate-producing bacteria in inflammatory bowel disease. Journal of Clinical Microbiology 52: 398–406. DOI 10.1128/JCM.01500-13. [Google Scholar] [CrossRef]
Wang Y, Kasper LH (2014). The role of microbiome in central nervous system disorders. Brain, Behavior, and Immunity 38: 1–12. DOI 10.1016/j.bbi.2013.12.015. [Google Scholar] [CrossRef]
Weir A, Westerhoff P, Fabricius L, Hristovski K, von Goetz N (2012). Titanium dioxide nanoparticles in food and personal care products. Environmental Science & Technology 46: 2242–2250. DOI 10.1021/es204168d. [Google Scholar] [CrossRef]
Wijnhoven SW, Peijnenburg WJ, Herberts CA, Hagens WI, Oomen AG, Heugens EH, Roszek B, Bisschops J, Gosens I, van de Meent D (2009). Nano-silver-a review of available data and knowledge gaps in human and environmental risk assessment. Nanotoxicology 3: 109–138. DOI 10.1080/17435390902725914. [Google Scholar] [CrossRef]
Wilding LA, Bassis CM, Walacavage K, Hashway S, Leroueil PR, Morishita M, Maynard AD, Philbert MA, Bergin IL (2016). Repeated dose (28-day) administration of silver nanoparticles of varied size and coating does not significantly alter the indigenous murine gut microbiome. Nanotoxicology 10: 513–520. DOI 10.3109/17435390.2015.1078854. [Google Scholar] [CrossRef]
Willing BP, Dicksved J, Halfvarson J, Andersson AF, Lucio M, Zheng Z, Järnerot G, Tysk C, Jansson JK, Engstrand L (2010). A pyrosequencing study in twins shows that gastrointestinal microbial profiles vary with inflammatory bowel disease phenotypes. Gastroenterology 139: 1844–1854, e1841. DOI 10.1053/j.gastro.2010.08.049. [Google Scholar] [CrossRef]
Wu H, Yin JJ, Wamer WG, Zeng M, Lo YM (2014). Reactive oxygen species-related activities of nano-iron metal and nano-iron oxides. Journal of Food and Drug Analysis 22: 86–94. DOI 10.1016/j.jfda.2014.01.007. [Google Scholar] [CrossRef]
Xia T, Lai W, Han M, Han M, Ma X, Zhang L (2017). Dietary ZnO nanoparticles alters intestinal microbiota and inflammation response in weaned piglets. Oncotarget 8: 64878–64891. DOI 10.18632/oncotarget.17612. [Google Scholar] [CrossRef]
Yan G, Huang Y, Bu Q, Lv L, Deng P, Zhou J, Wang Y, Yang Y, Liu Q, Cen X (2012). Zinc oxide nanoparticles cause nephrotoxicity and kidney metabolism alterations in rats. Journal of Environmental Science and Health, Part A 47: 577–588. DOI 10.1080/10934529.2012.650576. [Google Scholar] [CrossRef]
Zeevi D, Korem T, Godneva A, Bar N, Kurilshikov A, Lotan-Pompan M, Weinberger A, Fu J, Wijmenga C, Zhernakova A (2019). Structural variation in the gut microbiome associates with host health. Nature 56: 43–48. DOI 10.1038/s41586-019-1065-y. [Google Scholar] [CrossRef]
Zeng M, Inohara N, Nuñez G (2017). Mechanisms of inflammation-driven bacterial dysbiosis in the gut. Mucosal Immunology 10: 18–26. DOI 10.1038/mi.2016.75. [Google Scholar] [CrossRef]
Zhai Q, Li T, Yu L, Xiao Y, Feng S, Wu J, Zhao J, Zhang H, Chen W (2017). Effects of subchronic oral toxic metal exposure on the intestinal microbiota of mice. Science Bulletin 62: 831–840. DOI 10.1016/j.scib.2017.01.031. [Google Scholar] [CrossRef]
Zhang Q, Zhang K, Xu D, Yang G, Huang H, Nie F, Liu C, Yang S (2014). CuO nanostructures: Synthesis, characterization, growth mechanisms, fundamental properties, and applications. Progress in Materials Science 60: 208–337. DOI 10.1016/j.pmatsci.2013.09.003. [Google Scholar] [CrossRef]
Cite This Article
 Copyright © 2023 The Author(s). Published by Tech Science Press.
Copyright © 2023 The Author(s). Published by Tech Science Press.This work is licensed under a Creative Commons Attribution 4.0 International License , which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.


 Submit a Paper
Submit a Paper Propose a Special lssue
Propose a Special lssue View Full Text
View Full Text Download PDF
Download PDF Downloads
Downloads
 Citation Tools
Citation Tools